Research - International Journal of Medical Research & Health Sciences ( 2021) Volume 10, Issue 8
An Ensemble Model for Detecting Coronavirus disease-19 from Chest X-ray Images
Marwan A. Albahar1*, Mohammed I. Thanoon1 and Abdulaziz A. Albahr22College of Applied Medical Sciences, Basic Science, King Saud bin Abdulaziz University for Health Sciences, Alhasa, Saudi Arabia
Marwan A. Albahar, College of Computers at Al-Leith, Umm Al Qura University, Mecca, Saudi Arabia, Email: mabahar@uqu.edu.sa
Received: 28-Jul-2021 Accepted Date: Aug 24, 2021 ; Published: 31-Aug-2021
Abstract
In current times, novel Coronavirus Disease-19 (COVID-19) has become one of the world’s biggest challenges. The Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) which causes this disease results in high mortality and morbidity rates globally. It has been revealed from the study that patients infected with COVID-19 exhibit different radiographic visual characteristics along with dyspnea, fatigue, dry cough, fever, etc. In this account, Chest X-Ray (CXR) is considered as one of the most significant, non-invasive clinical methods that assist in detecting the visual responses that are related to SARS-CoV-2 infection. However, the radiologist’s and experts’ minimal availability and experts’ ability to interpret the CXR images and subtle images of the disease serve as the prominent hurdles in manual diagnosis. This study presents an automatic coronavirus disease-19 screening ensemble model that utilizes radiomic texture descriptors extracted from CXR images to identify normal, infected, and suspected coronavirus disease-19 patients. The ensemble model utilizes a majority vote-based classifier ensemble of two benchmark supervised classification algorithms. The training and testing of the proposed model are performed using the two datasets. The ensemble model can significantly improve prediction performance with 94% accuracy, while the precision and recall rate of detecting coronavirus disease-19 are both 93%.
Keywords
Artificial intelligence, Coronavirus disease 2019, Deep learning, Prognosis, X-Ray images
Introduction
Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) induced ailment named coronavirus disease-19 is a contagious disease initially detected in Wuhan City, Hubei Province, China, in December 2019. The disease first emerged in the seafood wholesale market and rapidly spread across the globe [1]. This infectious disease was declared a pandemic by the World Health Organization (WHO) on 11th March 2020 [2]. As of 20th September 2020, this life-threatening virus has harmed millions of people. To date, there have been approximately 30,675,675 confirmed coronavirus disease-19 patients and around 954,417 casualties [2]. To limit the spread of this lethal infection, all affected countries are making all possible efforts, like encouraging citizens to maintain social distance, practicing hygiene, enhancing the screening process through multi-functional testing, seeking massive vaccination to reduce the infection rate, etc. In this account, the Reverse Transcriptase-Polymerase Chain Reaction (RT-PCR) is widely used as a modular diagnosis method. However, it has been established that it possesses certain constraints, like a delayed diagnosis of suspected patients as the testing procedure takes some time at the clinical laboratory [3]. Another limitation associated with the test includes false-negative results that significantly hinder the disease’s control and prevention [4]. To overcome the shortcomings and inadequacies of RT-PCR testing, researchers are striving to promote a reliable and fast diagnostic method for coronavirus disease-19 infection. It has been established that the Wuhan University Zhongnan hospital and the WHO have issued quick guides that suggest that, along with the clinical symptoms, chest imaging could also be considered for diagnosing and treating this infection [5,6]. Rubin, et al. have contributed by providing a prolific guideline for medical practitioners that instructs the utilization of Computed Tomography (CT) and chest radiography for screening and assessing the disease’s progression [7]. Though CT scans have high sensitivity, they also have few limitations, like high cost and high radiation dose requirements during the screening process that exposes children and women to the risk of radiation [8]. However, chest X-ray-based diagnosis is the optimal option for detecting and treating coronavirus disease-19 infection. Ng, et al. stated that the pulmonary infection caused by coronavirus disease-19 can be correctly detected by chest X-ray images [9]. Besides, concerning Artificial Intelligence (AI)-based disease recognition systems, healthcare practitioners have also shed light on the significance of chest X-rays for exploring coronavirus disease-19 symptoms like opaque patterns in the lungs [10]. Furthermore, an ensemble machine learning paradigm that combines different classification algorithms for generating a single model is promising to obtain better predictive performance than could be obtained from any of the constituent learning algorithms alone. Additionally, an ensemble technique is deemed effective in making predictions as it produces a low error rate compared to a single model’s prediction. Also, they are famous for being faster and smaller than other architecture.
Related Works
Since there is a requirement for swiftly identifying the coronavirus disease-19 infections, the area of CNN-based AI systems is on a continuous rise. The main reason behind the boom in this specific area is that it would help speed up medical image analysis. Since chest X-ray screening is deemed the most effective procedure for detecting and diagnosing pneumonia and having the same symptoms, this procedure is also used for diagnosing coronavirus disease-19 infection [10]. This aspect has resulted in inspiring researchers to explore the capability of chest X-rays using different feature extraction methods, especially CNN-based approaches, for the detection of coronavirus disease-19; hence, playing a pivotal role in the efficient detection of infections and easing the burden of a healthcare system that is overwhelmed by the pandemic. A profound survey of the CNN technology application for detecting coronavirus disease-19 and automated lung segmentation was conducted in Shoeibi, Afshin, et al. [11]. In this account, a prime focus has been put on analyzing the use of Computed Tomography (CT) and X-rays. Additionally, Halgurd, et al., and Krizhevsky, et al. also conducted a test on modified pre-trained AlexNet and modified the CNN model using their CT scan dataset and chest X-ray [12,13]. The experiment gave an accuracy of approximately 98% via. modified pre-trained and 94.1% accuracy by utilizing modified CNN. Besides, Narin, et al. also tested three pre-trained models with ImageNet weights (such as Inception v3, ResNet50, and Inception-ResNet v2), while considering two image types-normal images and coronavirus disease-19. It resulted in giving the highest accuracy, i.e., approximately 98% [14-18].
Moreover, Wang, et al. introduced an entirely new CNN framework named COVID-Net and the large chest X-ray benchmark dataset “COVIDx” [19]. The authors got the highest accuracy of 93.3 while studying how COVIDNet utilizes the interpretability method to predict infection. Apostolopoulos and Mpesiana trained advanced CNN architectures (like MobileNetV2, VGG19, Xception, Inception, and Inception-ResNet v2) through transfer learning on ImageNet [16,18,20-23]. Moreover, different neural network architectures were also utilized. The fine-tuned models’ results depicted the proof-of-principle for utilizing CNN along with transfer learning, specifically for extracting radiological features. Minaee, et al. used a dataset of five thousand chest x-rays from publicly available datasets while using the subset of their benchmark for developing a model by fine-tuning the renowned pre-trained CNNs (like DenseNet121, SqueezeNet, ResNet50, and ResNet18) [17,24-26]. The evaluation of the proposed model was conducted using the remaining images that gave promising results in specificity and sensitivity. Eduardo, et al. proposed a deep learning framework that primarily extended the Efficient Net series introduced by Tan and Le, which is popular for its fewer computational steps and exceptional prediction performance [27,28]. The findings of the experiment gave notable classification performance, particularly in the cases of coronavirus disease-19 infections. Moreover, another method was proposed by Farooq, et al. named COVID-ResNet that makes use of the 3-step technique [29]. These included a gradual adjustment of the image size, automatic selection of the learning rate, and the fine-tuning of the pre-trained ResNet50 architecture to improve the model’s performance. Furthermore, DarkCovidNet, a CNN model, was proposed by Ozturk, et al. for automatically detecting coronavirus disease-19 through chest X-ray images [30]. In this study, the suggested method conducted two types of classifications. The first one was a binary classification (i.e., no-findings and COVID-19); however, the second classification was for multi-class (i.e., pneumonia, no-findings, and COVID-19). Ozturk, et al. provided an intuitive explanation using a heat map that helps the radiologist find and detect the infected area on the x-ray. In the same context, Ucar, et al. also conducted a study and proposed a finetuned lightweight SqueezeNet that obtained the fine-tuned hyper-parameters through Bayesian optimization [31]. The findings revealed that the proposed network had high performance compared to other CNN networks suggested for the detection of coronavirus disease-19 infection. Moreover, Karim, et al. also carried out a study and suggested using an explainable CNN-based method while demonstrating class discriminating regions named Deep COVID Explainer for automatically detecting coronavirus disease-19 from chest x-ray images [32]. Another research conducted by Afshar, et al. has also made an immense contribution to developing coronavirus disease-19 detection systems. In this account, the researchers used Capsule Networks (CapsNets) based on the CNN architecture [33]. The study’s findings showed that the system is efficient in terms of its statistical performance and minimal trainable parameters. Another model named CoroNet was proposed by Asif, et al. that utilized the Xception architecture pre-trained on the ImageNet dataset [34]. The architecture was trained on two publicly available datasets and conducted two diverse classification performance measurements, i.e., four and three classes, as their accuracy was 89.6% and 95%, respectively. In addition to this, a CNN-based model named CovXNet was proposed by Mohammad, et al. that utilizes depth-wise dilated convolution [35]. Firstly, the model was trained with some non-COVID-19 pneumonia images while transferring the collected learning to additional fine-tuning layers, resulting in refining the detection process. Since X-rays had different resolutions, the stacking algorithm was utilized for accurate predictions. The study results showed the accuracy of CovXNet in the case of multi-class classification (it was approximately 90%). Nihad, et al. proposed a CNN-based modular architecture called PDCOVIDNet (dilated convolution-based coronavirus disease-19 detection network) [36]. The architecture constituted numerous blocks that assisted in detecting coronavirus disease-19 infections from the chest X-rays of the patients. The finding claimed that the model is more effective than the existing CNN architecture, specifically with the recall and precision of 96.59% and 96.58%, respectively.
The literature review shows that most of the methods predict based on a single model’s output instead of an ensemble. However, some of the methods are dependent on the ensemble [32,35,37]. It has also been found that the ensemble brings numerous benefits in terms of minimizing prediction errors, hence categorizing it as the most versatile model. For example, a study conducted by Chowdhury, et al. presented an ensemble of CNN-based on EfficientNet for detecting coronavirus disease-19 infection [37]. In this account, a huge dataset of chest X-rays was used. During the study, the open-access X-ray collection was improved and transferred to ImageNet pre-trained weights for EfficientNet. Next, the top layers were fine-tuned, followed by an ensemble of model snapshots for classifying the chest X-ray matching. Though one of the studies, specifically Karim, et al., utilized ensembles of heterogeneous models, i.e., DenseNet161, ResNet18, and VGG19 [32]. However, the approach possessed few constraints. In particular, each model has to go through the training of many parameters, and each needs a separate training session. Moreover, Mahmud, et al. used the method of performing an ensemble on a single model [35]. However, it utilized different image resolutions. For each resolution, it produces a distinct model while stacking it for making predictions. This is deemed to be a lengthy procedure and requires computational overhead. Contrarily, Wang, et al. proposed and tested an advanced custom CNN architecture (COVID-Net) using the huge coronavirus disease-19 database [19]. However, because of several parameters, the model’s computation overhead was found to be exceptionally high. Therefore, these facts motivate us to involve and utilize the ensemble model for detecting coronavirus disease-19 from CXR Images. We use ensembles of machine learning algorithms to address the problems mentioned above, which leads to better accuracy and enhanced performance by combining homogeneous or heterogeneous classifiers with the coronavirus disease-19. The designed architecture commences with the scale of images, then optimizes and fine-tunes the models. After that, we extracted X-ray images’ features to be integrated into an ensemble voting to make predictions. The key contributions of the current research are listed as follows:
• We propose an ensemble model-based architecture that includes random forest and K-Nearest Neighbor algorithms. The rationale behind using these algorithms for ensemble learning is their high performance in detecting coronavirus disease-19
• Since it is assumed that multiple radiologists’ decisions are taken into consideration during the final predictions, the ensemble is used in the proposed model for making correct predictions, hence making a fair and credible evaluation
• Lastly, the research appraises the architecture with other advanced architecture by reviewing different studies, highlighting the suggested architecture’s efficacy in detecting coronavirus disease-19
Methodology
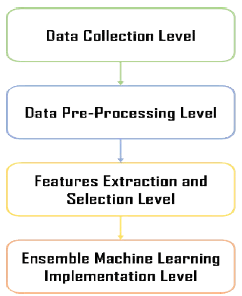
The present section briefly discusses the methodological approach that was used in the current research work. As with any machine learning model, the model’s architecture incorporates multiple levels. At the first level, we gathered the data from two public data sets. Then, the collected chest X-ray images are pre-processed at the second level. The third level of the proposed ensemble machine learning model is responsible for selecting features from the pre-processed chest X-ray images, while the last level of functionality is the development and implementation of the ensemble machine learning paradigm. Lastly, the final model was empirically evaluated by using state-of-the-art approaches to highlight the effectiveness and efficacy of the model in coronavirus disease-19 detection. Figure 1 illustrates the proposed ensemble machine learning model.
Implementation
In this section, a discussion of each level implementation of the proposed ensemble machine learning model will be outlined.
Data Collection Level
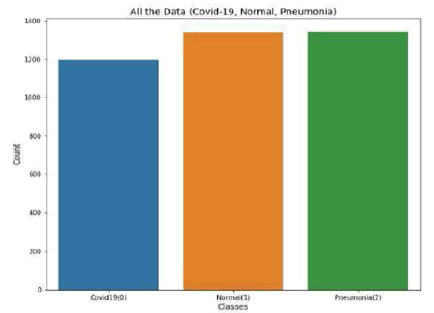
This study combined 2 public databases of coronavirus disease-19 chest X-rays. These included (1) COVID-19 Chest X-Ray Dataset Initiative and (2) COVID-19 Radiography Database [38,39]. The COVID-19 radiography database consists of 1341 normal images, 1143 coronavirus disease-19 positive images, and 1345 viral pneumonia images. Additionally, the COVID-19 Chest X-ray Dataset Initiative has 48 coronaviruses disease-19 positive images. The distribution of data based on the infection types from the two datasets is shown in Figure 2.
Data Pre-processing Level
The data was pre-processed to be prepared for the classification task. Images transformed the classes into the array during the first phase. Then, the image feature is converted into arrays. Finally, we normalize the features by dividing each feature value by 255 and arranging all feature values between the range of 0 and 1 from 0 to 255.
Features Selection Level
Feature selection can be understood as the most important process used in developing a machine learning model. The selection of adequate features helps minimize data redundancy while avoiding noisy data; hence, it enhances the model’s overall performance. We applied two techniques for feature extraction and selection: vector-based features and color histogram features. There are total features of 300 × 300=90000 and we selected 256 × 256=65536.
Ensemble Machine Learning Model Level
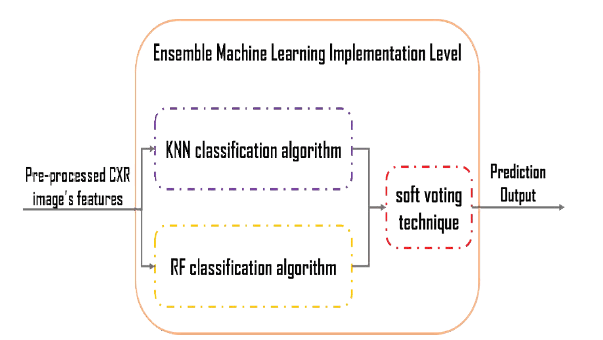
The ensemble machine learning paradigm is a combination of various classification algorithms used to generate a single model. In this research, we examined several classification algorithms, but we applied the two most accurate to the ensemble learning model: the KNN algorithm and the random forest algorithm. To implement the RF, two parameters are required to be set up. These include ‘n tree’ (the number of trees) and mtry (the number of features in each split). It has been claimed by several researchers that satisfactory results can be acquired with the default parameters [40-42]. However, as per Liaw and Wiener, the increased number of trees will offer more stable results, having variable importance [40]. Additionally, Breiman affirmed that the use of a higher number of trees, i.e., more than the required ones, is deemed to be unnecessary [43]. However, it is essential to note that it is not harmful to the model. Furthermore, Feng, et al. have also affirmed that accurate results can be achieved through RF, specifically with n tree=200 [44]. Concerning the mtry parameter, several studies have been carried out that utilize the default value mtry= √p. Here, p is regarded as the total number of predictor variables. Nevertheless, the present research has evaluated and tested the range of values for both parameters for finding an adequate RF model for classification. Moreover, the KNN approach is non-parametric in nature [45]. The basic theory involved in the operations of KNN is that in the calibration dataset, it searches for the group of ‘k’ samples, specifically the ones which are nearest to the unknown samples (i.e., it works on distance functions). These ‘k’ samples help determine the class (label) of unknown samples by estimating the average response variables [46]. Succinctly, it can be affirmed that ‘k’ plays a pivotal role in KNN’s performance. As per Zhang and Roy, ‘k’ is also regarded as KNN’s key tuning parameter [43]. It is important to note that the bootstrap procedure was used for determining the parameter ‘k.’ Therefore, the K parameter for the KNN algorithm and min_samples_leaf, min_samples_split, and random_state parameters for the random forest algorithm has to be tuned and adjusted. Fine-tuned hyper-parameters significantly impact the model’s performance because they directly govern the model’s training. Additionally, tuned parameters also assist in the production of highly accurate results when using RF and KNN. Each classifier has different tuning steps and tuned parameters. We tested a series of values for the tuning process with the optimal parameters determined based on each classifier’s highest classification accuracy. Consequently, we adjusted the hyper-parameter values as follows. For the KNN algorithm, we selected different values of k, but we got good results with k=2. For the Random Forest algorithm, after several trials, we concluded the following. The minimum samples of each tree were optimum when min_samples_leaf=10. For the minimum sample split of each tree, the best value occurs when min_samples_split=20. When random_state=3, we achieve reliable results. Furthermore, the model gets the extracted features of the CXR images as an input. Then, simultaneously, the nominated classification algorithms make a prediction. Finally, by using soft voting, we combined these two algorithms’ predictions. We set the voting classifier parameter as “soft” because we calculated the area’s probabilities under the curve plots. Figure 3 details the overall architecture of the proposed machine learning ensemble model.
Results
Due to the high variations between different executions, we evaluate the model using Accuracy, Precision, Recall, F1 Measure, Confusion Matrix, AUC Score, and ROC Curve in all the experiments. The used metrics are calculated as follows:
Accuracy=TP+TN/Total Predictions (1)
Precision=TP/TP+FP (2)
Recall=TP/TP+FN (3)
F1=Precision × Recall/Precision+Recall (4)
When calculating the accuracy matrices, the KNN and RF algorithms give 90%. However, using the proposed ensemble method, the accuracy rises to 94% as shown in Table 1.
| Model | Accuracy |
|---|---|
| KNN Algorithm | 0.90 |
| Random Forest Algorithm | 0.90 |
| Ensemble Learning Algorithm | 0.94 |
By making an allowance for each class perdition separately, the results are enhanced for all three considered algorithms. The reason behind that is because they possess more training weights as compared to the COVID-19 infection cases. The Precision, Recall, and F1-score metrics for the KNN, RF, and the proposed ensemble model are shown in Table 2, Table 3, and Table 4, respectively.
| Class | Precision | Recall | F1-score |
|---|---|---|---|
| 0 | 0.95 | 0.96 | 0.95 |
| 1 | 0.92 | 0.85 | 0.88 |
| 2 | 0.85 | 0.91 | 0.88 |
| Class | Precision | Recall | F1-score |
|---|---|---|---|
| 0 | 0.93 | 0.92 | 0.92 |
| 1 | 0.91 | 0.90 | 0.90 |
| 2 | 0.86 | 0.88 | 0.87 |
| Class | Precision | Recall | F1-score |
|---|---|---|---|
| 0 | 0.96 | 0.96 | 0.96 |
| 1 | 0.94 | 0.90 | 0.92 |
| 2 | 0.89 | 0.93 | 0.91 |
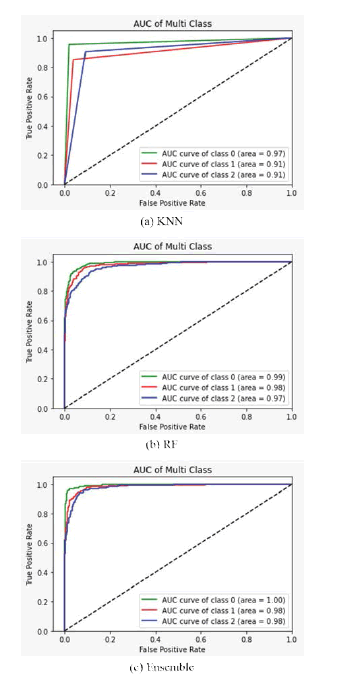
More deeply, using Accuracy, Precision, Recall, and F1-score is often difficult to measure the model’s performance. Thus, we need to look at the ROC curve, which allows a false positive rate since it plots the true positive rate against a false positive rate. In Figure 4, ROC curves display the micro and macro average and show robust AUC scores for all classes, showing the proposed model’s stable predictions and class-wise AUC scores achieved with our ensemble model. We achieved 1.00, 0.98, and 0.98 AUC scores for COVID-19, normal and viral pneumonia, respectively, in the ROC curves. Notably, the AUC of COVID-19 is significantly higher than in other classes.
Discussion
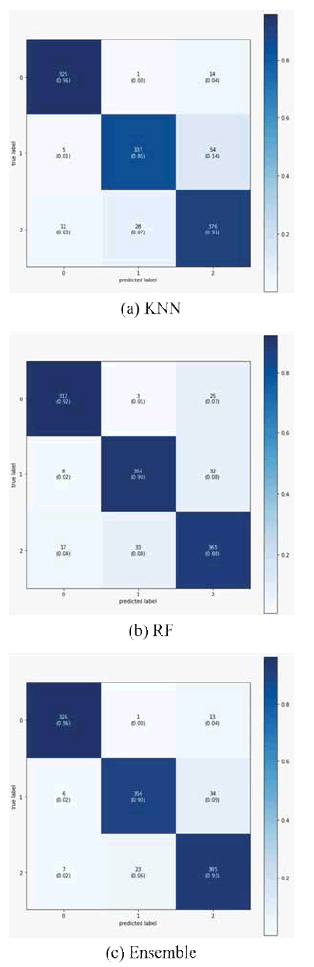
Figure 5 shows the confusion matrices for all evaluated models. It is clear that out of the 340 test images, 14 images were classified incorrectly. The performance of the KNN and ensemble models is similar to the COVID-19 image classification task. However, KNN and RF decline in other classes, while they increase slightly in the ensemble model. This behavior’s main reason is that COVID-19 infection is a special and relatively new type of viral pneumonia with slightly common features, resulting in misleading the models.
The findings show that KNN models possess good recall when classifying viral pneumonia and COVID-19. On the other hand, it presents declining trends in the prediction of normal cases. Besides, RF depicts the capability to detect normal images, but when it comes to detecting viral pneumonia, the performance declines compared to other models. Based on these findings, it can be established that the presented ensemble machine learning model is highly efficient at COVID-19 infection detection from chest X-ray images. Therefore, we believe that the proposed model focuses on discriminating features that can help distinguish between other types (e.g., normal and viral pneumonia). Table 5 shows the comparison between the proposed model and other models for detecting COVID-19 using chest X-rays, and we have seen that the proposed model is better than other models.
| Method | Total chest X-ray | Precision (%) (COVID-19) | Recall (%) (COVID-19) | Accuracy (%) |
|---|---|---|---|---|
| COVID-Net | 573 COVID-19 | 99.0 | 95.0 | 94.5 |
| 8,066 Normal | ||||
| 5559 Pneumonia | ||||
| EfficientNet-B3 [30] (Flat Classification) | 183 COVID-19 | 100 | 96.8 | 93.9 |
| 8,066 Normal | ||||
| 5,521 Pneumonia | ||||
| EfficientNet-B3 [30] (Hierarchical Classification) | 183 COVID-19 | 100 | 80.6 | 93.5 |
| 8,066 Normal | ||||
| 5,521 Pneumonia | ||||
| CovXNet [40] | 305 COVID-19 | 88.5 | 90.3 | 89.6 |
| 305 Viral Pneumonia | ||||
| 305 Bacterial Pneumonia | ||||
| 305 COVID-19 | 90.8 | 89.9 | 90.3 | |
| 305 Viral Pneumonia | ||||
| 305 Bacterial Pneumonia | ||||
| +305 Normal | ||||
| Proposed | 1198 COVID-19 | 96 | 96 | 94.0 |
| 1341 Normal | ||||
| 1345 Pneumonia |
Some previous models (namely EfficientNet-B3 and COVID-Net) used ImageNet weights and the COVIDx dataset [19,27]. CovXNet used an ensemble method and a transfer learning scheme from non-COVID chest X-rays while retaining training and testing datasets other than COVIDx [35]. COVID-Net demonstrated comparable results to our proposed system. However, due to the training of several parameters, the complexity of COVID-Net is high. Thus, the computational overhead limits its applicability. When comparing all ensemble models’ results, we observed that our ensemble showed satisfactory results in classifying COVID-19 with fewer parameters and computational overhead. For the cases associated with wrongly classified COVID-19 infection, it was found that seven (7) images of viral pneumonia were classified as COVID-19 infection. This could be due to the infections’ overlapping or similar characteristics, as both of the infections cause harm to the respiratory system/lungs. Undoubtedly, these overlapping characteristics cause confusion that demands clinical practitioners to thoroughly review the findings. Likewise, the proposed system also got confused in some stances, specifically during viral and normal pneumonia detection. One of the factors that can be associated with the progressive variation is the radiological manifestations in this account. For instance, viral pneumonia’s true image was classified as a normal infection as it is quite unrealistic to predict the infection at the early stages.
Conclusion
In this paper, we proposed an ensemble model for the identification of COVID-19 from chest X-ray images. As we have observed, the ensemble model can effectively capture COVID-19 features, so it has excellent classification performance compared to some well-known methods. The datasets used in the experiment have a good number of COVID-19 images. Our experimental evaluation shows that the ensemble model outperforms other models, with a precision and recall of 93% and 93%, respectively. The ensemble model demonstrates its potential through other performance metrics such as the weighted average of precision, recall, F1 scores, and overall model accuracy. For future research, diverse datasets of COVID-19 cases can be integrated and explored to increase the proposed model’s resilience. Additionally, a mobile application will also be developed that will incorporate a human-explainable function. It is anticipated that the mobile application will assist in effectively screening the COVID-19 infection; hence, assist in controlling this pandemic while preventing other potential pandemics that might occur in the future. Additionally, the model will be further extended to analyze the patients’ short-term historical chest X-ray patterns. This would eventually help in predicting whether the infection will threaten the life of the patient or not. Since studying the radiological markers of COVID-19 is an active research area, several aspects require further exploration. Considering this factor, future research will further examine the visualization techniques used to critically interpret the unique characteristics and features of COVID-19.
Declarations
Conflicts of Interest
The authors declared no potential conflicts of interest concerning the research, authorship, and/or publication of this article.
Ethical Approval
This article does not contain any studies with human participants performed by any of the authors.
Data Availability
Datasets used to support the findings of this study are included in the article.
Funding
This research work has received no funds.
References
- WHO. "Covid-2019 situation reports" World Health Organization, 2019.
- WHO. "Covid-19 pandemic" World Health Organization, 2019.
- Zheng, Chuansheng, et al. "Deep learning-based detection for COVID-19 from chest CT using weak label." MedRxiv, 2020.
- Fang, Yicheng, et al. "Sensitivity of chest CT for COVID-19: Comparison to RT-PCR." Radiology, Vol. 296, No. 2, 2020, pp. E115-17.
- World Health Organization. "Use of chest imaging in COVID-19: a rapid advice guide, 11 June 2020." No. WHO/2019-nCoV/Clinical/Radiology_imaging/2020.1, World Health Organization, 2020.
- Jin, Ying-Hui, et al. "A rapid advice guideline for the diagnosis and treatment of 2019 novel Coronavirus (2019-nCoV) infected pneumonia (standard version)." Military Medical Research, Vol. 7, No. 1, 2020, pp. 1-23.
- Rubin, Geoffrey D., et al. "The role of chest imaging in patient management during the COVID-19 pandemic: A multinational consensus statement from the Fleischner Society." Radiology, Vol. 296, No. 1, 2020, pp. 172-80.
- Davies, H. E., C. G. Wathen, and F. V. Gleeson. "The risks of radiation exposure related to diagnostic imaging and how to minimise them." BMJ, Vol. 342, 2011.
- Ng, Ming-Yen, et al. "Imaging profile of the COVID-19 infection: Radiologic findings and literature review." Radiology: Cardiothoracic Imaging, Vol. 2, No. 1, 2020, p. e200034.
- Rajpurkar, Pranav, et al. "Chexnet: Radiologist-level pneumonia detection on chest x-rays with deep learning." arXiv preprint arXiv:1711.05225, 2017.
- Shoeibi, Afshin, et al. "Automated detection and forecasting of covid-19 using deep learning techniques: A review." arXiv preprint arXiv:2007.10785, 2020.
- Maghdid, Halgurd S., et al. "Diagnosing COVID-19 pneumonia from X-ray and CT images using deep learning and transfer learning algorithms." Multimodal Image Exploitation and Learning, Vol. 11734, 2021.
- Krizhevsky, Alex, Ilya Sutskever, and Geoffrey E. Hinton. "Imagenet classification with deep convolutional neural networks." Advances in Neural Information Processing Systems, Vol. 25, 2012, pp. 1097-105.
- Ozturk, Tulin, et al. "Automated detection of COVID-19 cases using deep neural networks with X-ray images." Computers in Biology and Medicine, Vol. 121, 2020, p. 103792.
- Deng, Jia, et al. "Imagenet: A large-scale hierarchical image database." 2009 IEEE Conference on Computer Vision and Pattern Recognition, IEEE, 2009, pp. 248-55.
- Szegedy, Christian, et al. "Rethinking the inception architecture for computer vision." Proceedings of the IEEE Conference on Computer Vision znd Pattern Recognition, 2016, pp. 2818-26.
- He, Kaiming, et al. "Deep residual learning for image recognition." Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2016, pp. 770-78.
- Szegedy, Christian, et al. "Inception-v4, inception-resnet and the impact of residual connections on learning." Thirty-first AAAI Conference on Artificial Intelligence, 2017, pp. 4278-84.
- Wang, Linda, Zhong Qiu Lin, and Alexander Wong. "Covid-net: A tailored deep convolutional neural network design for detection of covid-19 cases from chest x-ray images." Scientific Reports, Vol. 10, No. 1, 2020, pp. 1-12.
- Apostolopoulos, Ioannis D., and Tzani A. Mpesiana. "Covid-19: Automatic detection from x-ray images utilizing transfer learning with convolutional neural networks." Physical and Engineering Sciences in Medicine, Vol. 43, No. 2, 2020, pp. 635-40.
- Sandler, Mark, et al. "Mobilenetv2: Inverted residuals and linear bottlenecks." Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2018 pp. 4510-20.
- Simonyan, Karen, and Andrew Zisserman. "Very deep convolutional networks for large-scale image recognition." arXiv preprint arXiv:1409.1556, 2014.
- Chollet, François. "Xception: Deep learning with depthwise separable convolutions." Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2017, pp. 1251-58.
- Minaee, Shervin, et al. "Deep-covid: Predicting covid-19 from chest x-ray images using deep transfer learning." Medical Image Analysis, Vol. 65, 2020, p. 101794.
- Huang, Gao, et al. "Densely connected convolutional networks." Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2017, pp. 4700-08.
- Iandola, Forrest N., et al. "SqueezeNet: AlexNet-level accuracy with 50x fewer parameters and<0.5 MB model size." arXiv preprint arXiv:1602.07360, 2016.
- Luz, Eduardo, et al. "Towards an effective and efficient deep learning model for COVID-19 patterns detection in X-ray images." Research on Biomedical Engineering, 2021, pp. 1-14.
- Tan, Mingxing, and Quoc Le. "Efficientnet: Rethinking model scaling for convolutional neural networks." International Conference on Machine Learning, PMLR, 2019, pp. 6105-14.
- Farooq, Muhammad, and Abdul Hafeez. "Covid-resnet: A deep learning framework for screening of covid19 from radiographs." arXiv preprint arXiv:2003.14395, 2020.
- Ozturk, Tulin, et al. "Automated detection of COVID-19 cases using deep neural networks with X-ray images." Computers in Biology and Medicine, Vol. 121, 2020, p. 103792.
- Ucar, Ferhat, and Deniz Korkmaz. "COVIDiagnosis-Net: Deep Bayes-SqueezeNet based diagnosis of the coronavirus disease 2019 (COVID-19) from X-ray images." Medical Hypotheses, Vol. 140, 2020, p. 109761.
- Karim, Md, et al. "DeepCOVIDExplainer: Explainable COVID-19 diagnosis based on chest X-ray images." arXiv preprint arXiv:2004.04582, 2020.
- Afshar, Parnian, et al. "Covid-caps: A capsule network-based framework for identification of covid-19 cases from x-ray images." Pattern Recognition Letters, Vol. 138, 2020, pp. 638-43.
- Khan, Asif Iqbal, Junaid Latief Shah, and Mohammad Mudasir Bhat. "CoroNet: A deep neural network for detection and diagnosis of COVID-19 from chest x-ray images." Computer Methods and Programs in Biomedicine, Vol. 196, 2020, p. 105581.
- Mahmud, Tanvir, Md Awsafur Rahman, and Shaikh Anowarul Fattah. "CovXNet: A multi-dilation convolutional neural network for automatic COVID-19 and other pneumonia detection from chest X-ray images with transferable multi-receptive feature optimization." Computers in Biology and Medicine, Vol. 122, 2020, p. 103869.
- Chowdhury, Nihad K., Md Muhtadir Rahman, and Muhammad Ashad Kabir. "PDCOVIDNet: A parallel-dilated convolutional neural network architecture for detecting COVID-19 from chest X-ray images." Health Information Science and Systems, Vol. 8, No. 1, 2020, pp. 1-14.
- Chowdhury, Nihad Karim, et al. "ECOVNet: An ensemble of deep convolutional neural networks based on efficientnet to detect COVID-19 from chest X-rays." arXiv preprint arXiv:2009.11850, 2020.
- Kaggle, Rahman T. "Covid-19 radiography database." 2020.
- Chung, A. "Figure 1 COVID-19 chest x-ray data initiative; 2020."
- Liaw, Andy, and Matthew Wiener. "Classification and regression by randomForest." R News, Vol. 2, No. 3, 2002, pp. 18-22.
- Immitzer, Markus, Clement Atzberger, and Tatjana Koukal. "Tree species classification with random forest using very high spatial resolution 8-band WorldView-2 satellite data." Remote Sensing, Vol. 4, No. 9, 2012, pp. 2661-93.
- Zhang, Hankui K., and David P. Roy. "Using the 500 m MODIS land cover product to derive a consistent continental scale 30 m Landsat land cover classification." Remote Sensing of Environment, Vol. 197, 2017, pp. 15-34.
- Breiman, Leo. "Random forests." Machine Learning, Vol. 45, No. 1, 2001, pp. 5-32.
- Feng, Quanlong, Jiantao Liu, and Jianhua Gong. "UAV remote sensing for urban vegetation mapping using random forest and texture analysis." Remote Sensing, Vol. 7, No. 1, 2015, pp. 1074-94.
- Franco-Lopez, Hector, Alan R. Ek, and Marvin E. Bauer. "Estimation and mapping of forest stand density, volume, and cover type using the k-nearest neighbors method." Remote Sensing of Environment, Vol. 77, No. 3, 2001, pp. 251-74.
- Akbulut, Yaman, et al. "Ns-k-nn: Neutrosophic set-based k-nearest neighbors classifier." Symmetry, Vol. 9, No. 9, 2017, p. 179.