Research - International Journal of Medical Research & Health Sciences ( 2021) Volume 10, Issue 2
Detection of Toxins and Antibiotic Resistance Genes Profile among Methicillin-Resistant Staphylococcus aureus (MRSA) Isolates and Their Types of Infection in a Tertiary Hospital in Malaysia
Norhafez Senon1, Hassanain Al-Talib2* and Ariza Adnan22Medical Microbiology and Parasitology Department, Faculty of Medicine, Universiti Teknologi MARA (UiTM), Sungai Buloh, Selangor, 47000, Malaysia
Hassanain Al-Talib, Medical Microbiology and Parasitology Department, Faculty of Medicine, Universiti Teknologi MARA (UiTM), Sungai Buloh, Selangor, 47000, Malaysia, Tel: +60 3-5544 2000, Email: hassanainiy@yahoo.com
Received: 30-Dec-2020 Accepted Date: Feb 11, 2021 ; Published: 18-Feb-2021
Abstract
Objective: This study aimed to determine the molecular characteristics of virulence and antibiotics resistance genes profile of MRSA isolates and to compare between the conventional and molecular patterns of antibiotic resistance and determine the association between the MRSA virulence genes with their types of infection. Methods: A total of 60 MRSA isolates were collected from the Microbiology Laboratory of Selayang Hospital, a tertiary hospital in Malaysia. Antimicrobial susceptibility tests were performed by the disc diffusion method. The virulence and the antibiotic resistance genes were determined by PCR. Results: Among the sixty clinical isolates, there were six types of MRSA infections including cellulitis (30%), diabetic foot ulcer (28.3%), necrotizing fasciitis (13.3%), osteomyelitis (15%), catheter-related bloodstream infection (10%) and pneumonia (3.3%). Overall, lukS genes were detected in 60% followed by staphylococcal enterotoxins A (sea) gene (45%), toxic shock syndrome toxin (tsst) gene (43.3%), and 8.3% for each exfoliative toxin A (eta) and α-hemolysin (hla) genes. A significant association was found between sea and hla genes with all the types of infection. The sea genes were significantly associated with cellulitis, diabetic foot ulcer, and osteomyelitis. The detection of antibiotic resistance by the molecular and conventional methods was comparable. Conclusion: This study showed that there were multiple virulences and antibiotic-resistant genes involved in the pathogenicity of MRSA infections. Both sea and hla genes have a significant association with the various types of infections. Our outcomes showed an elevated rate of lukS gene among MRSA isolates. Vancomycin-resistance was not detected among the MRSA isolates.
Keywords
MRSA, Virulence genes, lukS, antibiotic resistance, sea
Introduction
Staphylococcus aureus is among the most common human pathogens and can cause numerous and serious infections. The highly virulent strain can cause severe diseases, which may lead to fatality. They can produce a wide range of toxins and exhibit resistance to single or multiple antibiotics. Most of these traits are located on mobile genetic elements of the bacterial genome [1]. Methicillin-resistant Staphylococcus aureus (MRSA) is a worldwide pathogen leading to hospital-acquired infections and is commonly resulted in significant mortality, morbidity, and increase of hospital stay cost burden [2]. Recently, MRSA infections have emerged in the community and also from livestock; however, they vary not only in their clinical aspects and molecular biology but also their antimicrobial susceptibility and treatment [3]. MRSA strains have evolved by acquiring a genetic element known as Staphylococcal Cassette Chromosome (SCC) mec, which contains the mobile mecA gene. Acquiring of the mecA gene codes for an altered Penicillin-Binding Protein (PBP) that has a lower affinity for binding β-lactams [4]. Risk factors associated with MRSA infection include prolonged hospitalization, previous antibiotic administration, and invasive procedures in addition to MRSA colonization [2]. A higher incidence of MRSA infection is also reported among healthcare workers who are dealing with patients infected with this pathogen [3]. MRSA can cause a variety of infections involving skin and subcutaneous tissues, to invasive infections like osteomyelitis, meningitis, pneumonia, lung abscess, and infective endocarditis [5].
A wide range of virulence factors, including toxins, immune-evasive surface factors, and enzymes that promote tissue invasion are produced by MRSA [6]. The virulence genes of interest in this study were collectively targeted for the first time in Malaysia including staphylococcal enterotoxin A gene (sea), toxic shock syndrome toxin gene (tsst), exfoliative toxin A gene (eta), Panton-Valentine leukocidin gene (lukS), and α-hemolysin gene (hla).
MRSA can produce a variety of Staphylococcal Enterotoxins (SEs). These toxins have a potent superantigenic role and are consisted of nine types (SEA to SEI). Few of these SEs have a vital role in epidemics of food poisoning and other infections that are septic-related. These SE proteins have a significant ability to resist heat and acid. Therefore, they may not be denatured by simple cooking of contaminated food [7].
Staphylococcal enterotoxins act as strong gastrointestinal toxins and as superantigens that spur nonâ??specific T-cell proliferation [8]. Though these are two distinct actions located on detaching domains of the proteins, there is a high association between these actions i.e., a loss of superantigen activity (due to genetic mutation) results in loss of enterotoxic activity also [9]. Another toxin produced by MRSA is the toxic shock syndrome toxin encoded by the tsst-1 gene. The tsst-1 gene encodes a 21.9 KDa extracellular toxin causing toxic shock syndrome. The tsst-1 triggers a T cell-dependent shock syndrome resulting in high lethality by stimulating the release of cytokines, including interleukin-6 and tumor necrosis factors [10].
Exfoliative toxins (ETA, ETB, and ETD) are produced by MRSA which act as proteases that hydrolyze desmoglein-1 (DG-1), an important keratinocyte help in cell-cell adhesion in the superficial epidermis. Hydrolysis of desmoglein caused dissociation of keratinocytes in humans a condition called staphylococcal scalded skin syndrome [11].
Panton-Valentine Leukocidin (PVL) is another important cytotoxin made by MRSA strains, which is encoded by two separate genes, lukS-PV and lukF-PV. The PVL is a synergohymenotropic toxin, and it acts through the synergistic activity of 2 non-associated secretory proteins, component S and component F that induces pores in the membranes of cells [12]. PVL production was initially linked to furuncles, cutaneous abscesses, and severe necrotic skin infections, though; severe cases of necrotizing hemorrhagic pneumonia and septicemia have also been reported. PVL genes are carried mainly by Community-Acquired MRSA (CA-MRSA). However, PVL genes are seldomly present in hospital isolates; hence it is known as a marker of CA-MRSA. Epidemiological records indicate that high severity of CAMRSA is reported with PVL genes [13].
Red blood cell lysis by MRSA is mainly mediated by the different hemolysins known as alpha, beta, delta toxins, and gamma hemolysins. The alpha-hemolysin is a pore-forming toxin encoded by the hla gene. α-hemolysin is the most characterized virulence factor of S. aureus that forms heptameric pores in host cell membranes, leading to lysis of the cell [14]. Also, α-hemolysin has been shown to affect innate immune effector cells, accelerate a hyper-inflammatory response characteristic of bacterial pneumonia, and disrupt epithelial and endothelial barriers. The hla expression is controlled by a complex regulatory network and its expression has been reported to be up-regulated during infection [15].
Mobile genetic elements carrying antibiotic resistance genes have been acquired by MRSA on multiple independent occasions and it reflects the strong selective pressures within a hospital environment [6]. Antibiotic resistance genes which were selected in this study included vancomycin-resistant gene (vanA), linezolid resistant gene (cfr), mupirocin resistant gene (mupA), rifampicin-resistant gene (rpoB), fusidic acid-resistant gene (fusA), and clindamycin resistant gene (ermA) represented the common genes in MRSA.
The high rate of antibiotic resistance and increasing multi-drug resistant bacteria is a promising challenge. Biocompatible nanostructures or nanocomposites such as reduced graphene oxide have been reported to exhibit strong antibacterial activity toward both Gram-positive and Gram-negative bacteria and make it an interesting candidate to incorporate into wound bandages to treat and/or prevent microbial infections [16,17].
In Malaysia, only a few studies had targeted MRSA virulence and antibiotics resistance characteristics and the infections they caused. Hence, this study was conducted to determine the virulence and antibiotics resistance profiles of the MRSA strains and the association with their types of infection.
Materials and Methods
This was a cross-sectional study that was carried out from March to June 2018 in Hospital Selayang, a tertiary, 800-bedded hospital in the suburb of Kuala Lumpur after approval was obtained from the Research and Ethical Committee. This study complies with the Declaration of Helsinki, Informed consent was obtained from the participants. A total of 60 MRSA isolates from clinical samples received at the microbiology laboratory of the hospital were included in the study. The clinical samples were from both adults and pediatric patients admitted to the hospital. Only one MRSA isolate per patient was included. The patients’ demography and types of infections were obtained from their medical records.
Conventional Methods
Identification of S. aureus was determined by standard microbiology practices based on colonial morphology, Gram stain, catalase test, and coagulase test. The Disc-diffusion agar method was used for the differentiation of MRSA from Methicillin-Sensitive S. aureus (MRSA) strains. The isolates were suspended in a broth medium of 0.5 McFarland standard concentrations and then grown in Muller Hinton agar. Cefoxitin (30 μg) discs were used as a control. All plates were incubated overnight at 37°C. The diameter of the inhibition zone around the discs was recorded and interpretation was in accordance with the Clinical Laboratory Standard Institute (CLSI) guideline with inhibition zone diameter ≤ 21 specified MRSA and inhibition zone diameter ≥ 22 suggested MSSA [18].
Conventional Identification of Antibiotic Resistance
All the isolates were tested for antibiotic susceptibility testing towards clindamycin (2 μg), linezolid (30 μg), mupirocin (5 μg), rifampicin (5 μg), fusidic acid (10 μg) using disc-diffusion agar method while vancomycin using the MIC E-test® strips (BioMérieux, Marcy l’Étoile, France). Both the methods were performed following the CLSI [18].
Method for the Identification of Virulence and Antibiotic Resistance Genes
DNA extraction: One bacterial colony was suspended in 200 μl of TE buffer (Tris-HCl [10 mM]: EDTA and boiled for 10 minutes then centrifuged for 5 minutes at 14,000 rpm at room temperature. The supernatant containing DNA (100 μl) was used for PCR assay.
PCR assay: All the 60 MRSA samples were tested for virulence (sea, tsst, eta, lukS, and hla) and antibiotic resistance (vanA, cfr, mupA, rpoB, ermA, and fusA) genes by PCR using previously described primers Table 1 [19-25].
| Oligonucleotide sequence (5’-3’) | Amplicon size, bp | Reference | ||
|---|---|---|---|---|
| sea | sea1 (F) | GGTTATCAATGTGCGGGTGG | 102 | [20] |
| sea2 (R) | CGGCACTTTTTTCTCTTCGG | |||
| tsst | tsst1 (F) | ACCCCTGTTCCCTTATCATC | 326 | [20] |
| tsst2 (R) | TTTTCAGTATTTGTAACGCC | |||
| eta | eta1 (F) | GCAGGTGTTGATTTAGCATT | 93 | [20] |
| eta2 (R) | AGATGTCCCTATTTTTGCTG | |||
| lukS | pvl1 (F) | ATCATTAGGTAAAATGTCTGGACATGATCCA | 443 | [20] |
| pvl2 (R) | GCATCAAGTGTATTGGATAGCAAAAGC | |||
| hla | hla1 (F) | CTGATTACTATCCAAGAAATTCGATTG | 209 | [20] |
| hla2 (R) | CTTTCCAGCCTACTTTTTTATCAGT | |||
| vanA | vanA1 (F) | ATGAATAGAATAAAAGTTGC | 1031 | [25] |
| vanA2 (R) | TCACCCCTTTAACGCTAATA | |||
| cfr | cfr1 (F) | TGAAGTATAAAGCAGGTTGGGAGTCA | 746 | [22] |
| cfr2 (R) | ACCATATAATTGACCACAAGCAGC | |||
| mupA | mup1 (F) | TATATTATGCGATGGAAGGTTGG | 456 | [21] |
| mup2 (R) | AATAAAATCAGCTGGAAAGTGTTG | |||
| rpoB | rpoB1 (F) | GTCGTTTACGTTCTGTAGGTG | 432 | [23] |
| rpoB2 (R) | TCAACTTTACGATATGGTGTTTC | |||
| ermA | ermA1 (F) | GTTCAAGAACAATCAATACAGAG | 421 | [19] |
| ermA2 (R) | GGATCAGGAAAAGGACATTTTAC | |||
| fusA | fusA1 (F) | CGCGGATCCTATCGTATTTATTCAGTAAT | 2100 | [24] |
| fusA2 (R) | AAGGATCCCTTGTATTTTAACCTAGGCTA | |||
Table 1: The oligonucleotide sequences and amplicon size of the virulence and antibiotic resistance genes in MRSA.
Standard strains known to be positive for the respective genes were used as positive controls (Table 2).
| Strains | Target genes |
|---|---|
| S. aureus ATCC 14458 | lukS |
| S. aureus N315 | hla, eta |
| S. aureus JCSC/4469 | tsst/sea |
| E. faecium BM4147 | vanA |
| S. aureus NRS 119 | cfr |
| S. aureus ATCC BAA1708 | mupA |
| S. aureus ATCC BAA44 | rpoB |
| S. aureus ATCC 700699 | fusA |
| S. aureus ATCC BAA977 | ermA |
Table 2 The control strains used in the detection of virulence and antibiotic resistance genes among MRSA isolates.
The optimal concentrations of primers for each gene 1 pmol were used in the multiplex PCR. The other PCR components used included 25 mM MgCl2, 10 × PCR buffer, 5 U Taq DNA polymerase, and 10 μM dNTPs. The PCR was accomplished using a Mastercycler Gradient Eppendorf, Hamburg, and Germany. The initial cycle of denaturation at 95°C for 4 min was followed by 30 cycles of denaturation at 95°C for 30 s, annealing for 30 s at range (59°C-45°C based on each primer), extension at 72°C for 1 min and a final extension at 72°C for 5 min. PCR products were mixed with 1 μL of loading buffer solution and neatly loaded in the agarose gel wells (1.5% plus Floro Red Acid Stain (1st Base, Singapore)) and electrophoresed at 75 V for 90 minutes.
Statistical Analysis
The data were analyzed by Statistical Package for Social Sciences (SPSS) software, version 25. Chi-square test was used to define the association among the two groups of variables with p<0.05 considered to be significant statistically.
Results
Among the 60 MRSA isolates 39 (65%) were isolated from Malay patients, followed by Chinese 11 (18.33%), Indian 9 (15%), and others 2 (1.7%). The results showed higher MRSA infections in male patients, with a sex ratio male to female of 1.3:1. The mean age of patients was 44.9 years and ranged from (1-72) years old. The MRSA infections were classified into six types of infections. Among the 60 clinical isolates; 18 (30%) were cellulitis 18 (30%), 17 (23.8%) were diabetic foot ulcer, 8 (13.3%) were necrotising fasciitis, 9 (15%) were osteomyelitis, 6 (10%) were catheter-related bloodstream infection and 2 (3.3%) were pneumonia. Antibiotic susceptibility testing revealed all the isolates were susceptible to linezolid, vancomycin, and mupirocin. Fifty-two (86.7%) of the isolates were susceptible to rifampicin, 54 (90.0%) susceptible to fusidic acid, and 57 (95.0%) susceptible to clindamycin (Table 3).
| Antibiotic (µg) | Conventional n (%) | Resistance gene | Molecular n (%) |
|---|---|---|---|
| Vancomycin * | 0 (0) | vanA | 0 (0) |
| Linezolid (30) | 0 (0) | cfr | 0 (0) |
| Mupirocin (5) | 0 (0) | mupA | 0 (0) |
| Rifampicin (5) | 8 (13.3) | rpoB | 5 (8.3) |
| Fusidic acid (10) | 6 (10) | fusA | 5 (8.3) |
| Clindamycin (2) | 3 (5) | ermA | 1(1.7) |
| *Determination of MIC by E-test | |||
Table 3: Comparison of antibiotic resistance of MRSA isolates by conventional and molecular methods.
The current study revealed five common virulence genes detected among the MRSA clinical isolates. The most common virulence gene was lukS (60%), followed by sea (45%), tsst (43.3%), eta (8.3%, and hla (8.3%). The sea gene was found in 27 (45%) isolates and 26 (43.3%) isolates possessed the tsst gene. The eta and hla genes were found in 5 (8.3%) isolates. The lukS was found in 36 (60%) isolates.
Table 4 shows the association between clinical diagnosis and the presence of virulence genes. The sea, tsst, and lukS genes were detected in a higher proportion in patients with MRSA infection. There is no significant difference between the various types of infections and the presence of tsst, eta, and lukS genes. The sea and lukS genes were detected in a higher proportion in patients with cellulitis, diabetic foot ulcer, and osteomyelitis. The sea genes were significantly associated with cellulitis, diabetic foot ulcer, and osteomyelitis (p<0.05). But the hla gene was not found in previous infections (p=0.006).
| Virulence genes | No. of positive isolates (n=60) | Virulence genes present in MRSA isolates according to infection types n (%) | ||||||
|---|---|---|---|---|---|---|---|---|
| Cellulitis (n=18) |
Diabetic foot ulcer (n=17) | Necrotising fasciitis (n=8) | Osteomyelitis (n=9) | CRBSI (n=6) |
Pneumonia (n=2) | p-value | ||
| sea | 27 (45) | 10 (55.6) | 8 (47.1) | 2 (25) | 6 (66.7) | 0 (0) | 1 (50) | 0.049 |
| tsst | 26 (43.3) | 8 (44.4) | 6 (35.3) | 3 (37.5) | 6 (66.7) | 3 (50) | 0 (0) | 0.423 |
| eta | 5 (8.3) | 1 (5.6) | 0 (0) | 1 (12.5) | 3 (33.3) | 0 (0) | 0 (0) | 0.101 |
| lukS | 36 (60) | 13 (72.2) | 8 (47.1) | 4 (50) | 7 (77.8) | 3 (50) | 1 (50) | 0.512 |
| hla | 5 (8.3) | 0 (0) | 0 (0) | 3 (37.5) | 0 (0) | 2 (33.3) | 0 (0) | 0.006 |
| CRBSI: Catheter-Related Blood Stream Infection | ||||||||
Table 4 Association between MRSA virulence genes and types of infection.
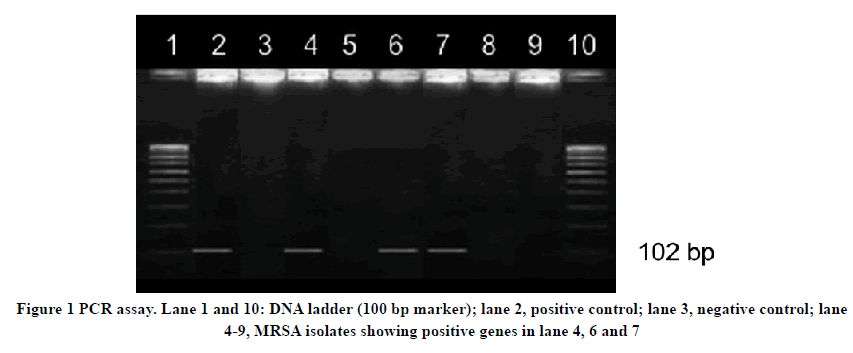
In this study, the virulence and antibiotic resistance genes were detected by PCR. All the detected genes were correctly matched to their base-pair according to the primers used from earlier studies. Figure 1 showed positive samples for sea genes from clinical MRSA isolates.
Discussion
In the study, the higher proportion of MRSA was among the Malay race (65%) which was due to the higher proportion of this ethnic group admitted to the Selayang Hospital. We had detected a range of antibiotic resistance and virulence genes among the MRSA isolates. Antibiotic-resistant testing of the MRSA isolates by molecular method showed resistance to rifampicin (8.3%), fusidic acid (8.3%), and clindamycin (1.7%). These results were in concordance with previous studies [19,23,24]. On the other hand, all MRSA isolates didn’t show a resistant gene band for vancomycin, linezolid, and mupirocin. These findings were similar to earlier studies by Thati, et al. in Hyderabad, Kehrenberg, et al. in Germany, and Akpaka et al. in Trinidad [22,25,26]. The presence of antibiotic resistance and virulence genes frequency differ globally and within the same country and even between different cities or hospitals. The difference in the percentage of these genes could be due to variations in geographical situations of each population or region or part of the hospital where the samplings were collected [20].
The study showed an insignificant discrepancy in the detection of antibiotic resistance patterns via disc diffusion and molecular methods. The Disc-diffusion method showed slightly higher resistance rates to rifampicin, fusidic acid, and clindamycin than the molecular method. This might be attributed to other mechanisms of antibiotic resistance that were not coded by the genes that were tested. Furthermore, it is known that there are other mechanisms of antibiotic resistance that contribute to the same antibiotic-resistant pattern such as decreased cell permeability, active efflux, enzymatic inactivation of the antibiotic, modification of the drug-receptor site, and synthesis of the resistant metabolic pathway. All the MRSA isolates were susceptible to vancomycin, linezolid, and mupirocin by both disc-diffusion and molecular methods. Similar findings were reported by current studies conducted in Hospital Tuanku Jaafar in Negeri Sembilan and University Malaya Medical Centre in Kuala Lumpur [27,28]. Fortunately, Vancomycin-Resistance S. aureus (VRSA) has not been documented in Malaysia thus far. However, many VRSA strains were already reported in neighboring countries from Indonesia and Thailand with a vancomycin MIC of more than ≥ 16 μg/mL [29,30]. Hence, we should constantly be vigilant of the VRSA strains as it is anticipated that they would eventually emerge in Malaysia.
Earlier studies have stated that the lukS and sea genes are the most common toxin genes in MRSA isolates [31]. Similarly, the lukS gene was the predominant gene (60%) detected in this study, which was also consistent with previous studies conducted in Malaysia which showed the lukS gene detection rate ranged between 3.8% to 11.1% in three different hospitals in the country [32-34]. The higher lukS generate in our study could be attributed to the types of infection as the MRSA infections in the patients were mostly acquired from the community whereby the presence of lukS gene is well documented or it might reflect a possible clonal expansion of those CA-MRSA into the hospital environment. Moreover, among the 60 clinical MRSA isolates 71.6% were of skin and soft tissue infections which were known to be highly associated with the presence of lukS gene [35]. The second highest virulent toxin which was detected from MRSA isolates was staphylococcal enterotoxin type A (sea) with 45%. Although, enterotoxin A is associated with the sporadic food-poisoning syndrome and foodborne outbreaks sea gene was the most common staphylococcal enterotoxin gene present among Malaysian MRSA strains which were isolated from invasive samples [36]. A previous study by Lim, et al. showed a significant increase in the prevalence of sea genes in 2008 strains compared with 2003 strains. These toxins can also cause toxic shock-like syndromes and have been implicated in several allergic and autoimmune diseases [8].
To the best of our knowledge, studies on tsst in Malaysia were scarce. Surprisingly, this study has demonstrated a high percentage of tsst gene among MRSA isolates i.e., 26 (46%) out of 60 isolates. The finding in this study was similar to a study conducted in Japan by Parsonnet and his colleagues; but lower than another study from Iran which showed 68% [37,38]. However, none of the MRSA isolates showed tsst gene in two earlier studies conducted in Kuala Lumpur and Terengganu [39,40]. The difference between the findings in this study and other literature may be due to a difference in geographic regions and also due to the differences in techniques as most of the previous studies evaluated the presence of anti-TSST antibodies.
Although the frequency of the tsst gene within Methicillin Sensitive Staphylococcus aureus strains (MSSA) is known to be higher, recently, it was reported that tsst expression was independent of the sensitivity of S. aureus to methicillin; also the number of MRSA strains harboring this gene has increased [37]. Several studies have also stated that staphylococcal toxins yielded vary considering the existence of different genotypes. Furthermore, it has been indicated that the virulence gene profiles of S. aureus in specific isolates may be influenced by the origins of their geographical place for example there was a high detection rate of tsst and sea genes in the United Kingdom and the United State of America [41,42]. An association between the MRSA virulence genes and the type of infections revealed no significant difference in the distribution of lukS among different types of MRSA infections. Although, hla has the lowest rate among other genes it showed a significant association with clinical MRSA cases (p-value<0.05). The result was not in concordance with the previous finding by Rossato, et al. who found the coexistence of hla genes in 87.6% among MRSA isolates obtained from hospitals in Porto Alegre, Brazil [43].
This study has several limitations including its small sample size. Also, the protein expression by virulence genes was not studied. Previous reports stated that the expression of drug resistance genes in MRSA strains might reduce the expression of virulence protein [44]. Another limitation is that this study used Chi-square in analysis, which does not provide the strength of the association between those two groups of variables.
Conclusion
The study revealed different characteristics of toxins and antibiotic resistance genes profiles between different MRSA infections. Both molecular and conventional methods for the detection of antibiotic resistance were comparable. All MRSA strains were sensitive to vancomycin, linezolid, and mupirocin. Both sea and hla genes have a significant association with the types of infection. Our outcomes showed a high rate of lukS gene among the MRSA isolates. Further studies needed to be carried out on the pathogenesis of these genes and their association with the types of MRSA infections.
Declarations
Acknowledgment
We are grateful to the staff from the Institute for Medical Molecular Biotechnology, Faculty of Medicine, (UiTM), and Microbiology laboratory of Selayang Hospital for their valuable support and assistance. This research was funded by 600-IRMI/DANA, 5/3/BESTARI (052/2017) grant from Universiti Teknologi MARA.
Conflict of Interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
References
- Malachowa, Natalia, and Frank R. DeLeo. "Mobile genetic elements of Staphylococcus aureus."Cellular and Molecular Life Sciences,Vol. 67, No. 18, 2010, pp. 3057-71.
- Al-Talib, Hassanain I., et al. "Methicillin-resistant Staphylococcus aureus nosocomial infection trends in Hospital Universiti Sains Malaysia during 2002-2007."Annals of Saudi Medicine,Vol. 30, No. 5, 2010, pp. 358-63.
- Siddiqui, Abdul H., and Janak Koirala. "Methicillin-Resistant Staphylococcus aureus (MRSA)."StatPearls [Internet],2019.
- Al-Talib, Hassanain, et al. "A pentaplex PCR assay for the rapid detection of methicillin-resistant Staphylococcus aureus and Panton-Valentine Leucocidin."BMC Microbiology,Vol. 9, No. 1, 2009, pp. 1-8.
- Contreras, Germán A., et al. "Empyema necessitans and acute osteomyelitis associated with community acquired methicillin resistant Staphylococcus aureus in an infant."Biomedica,Vol. 29, No. 4, 2009, pp. 506-12.
- Turner, Nicholas A., et al. "Methicillin-resistant Staphylococcus aureus: An overview of basic and clinical research."Nature Reviews Microbiology,Vol. 17, No. 4, 2019, pp. 203-18.
- Fri, Justine, et al. "Antibiotic resistance and virulence gene characteristics of Methicillin-Resistant Staphylococcus aureus (MRSA) isolated from healthy edible marine fish."International Journal of Microbiology,Vol. 2020, 2020.
- Ortega, Elena, et al. "Multiple roles of Staphylococcus aureus enterotoxins: Pathogenicity, superantigenic activity, and correlation to antibiotic resistance."Toxins,Vol. 2, No. 8, 2010, pp. 2117-31.
- Harris, T. O., et al. "Lack of complete correlation between emetic and T-cell-stimulatory activities of staphylococcal enterotoxins."Infection and Immunity,Vol. 61, No. 8, 1993, pp. 3175-83.
- Kim, Dokyun, et al. "Toxic shock syndrome toxin 1-producing methicillin-resistant Staphylococcus aureus of clonal complex 5, the New York/Japan epidemic clone, causing a high early-mortality rate in patients with bloodstream infections."Antimicrobial Agents and Chemotherapy,Vol. 63, No. 11, 2019, pp. e01362-19.
- Imanishi, Ichiro, et al. "Exfoliative toxin E, a new Staphylococcus aureus virulence factor with host-specific activity."Scientific Reports,Vol. 9, No. 1, 2019, pp. 1-12.
- Al-Talib, Hassanain, et al. "Fatal necrotizing pneumonia caused by Panton-Valentine leukocidin-producing hospital-acquired Staphylococcus aureus: A case report."Japanese Journal of Infectious Diseases,Vol. 64, No. 1, 2011, pp. 58-60.
- Bhatta, Dharm R., et al. "Association of Panton Valentine Leukocidin (PVL) genes with Methicillin Resistant Staphylococcus aureus (MRSA) in Western Nepal: A matter of concern for community infections (a hospital based prospective study)."BMC Infectious Diseases,Vol. 16, No. 1, 2016, pp. 1-6.
- Kong, Cin, Hui-min Neoh, and Sheila Nathan. "Targeting Staphylococcus aureus toxins: A potential form of anti-virulence therapy."Toxins,Vol. 8, No. 3, 2016, p. 72.
- Tabor, David E., et al. "Staphylococcus aureus alpha-toxin is conserved among diverse hospital respiratory isolates collected from a global surveillance study and is neutralized by monoclonal antibody MEDI4893."Antimicrobial Agents and Chemotherapy,Vol. 60, No. 9, 2016, pp. 5312-21.
- Di Giulio, Mara, et al. "Antimicrobial and antibiofilm efficacy of graphene oxide against chronic wound microorganisms."Antimicrobial Agents and Chemotherapy,Vol. 62, No. 7, 2018.
- Díez-Pascual, Ana María. "Antibacterial action of nanoparticle loaded nanocomposites based on graphene and its derivatives: A mini-review."International Journal of Molecular Sciences,Vol. 21, No. 10, 2020, p. 3563.
- Wayne, P. A. "Clinical and laboratory standards institute: Performance standards for antimicrobial susceptibility testing: 20th informational supplement."CLSI Document M100-S20,2010.
- Ghanbari, Fahimeh, et al. "Distribution of erm genes among Staphylococcus aureus isolates with inducible resistance to clindamycin in Isfahan, Iran."Advanced Biomedical Research,Vol. 5, 2016, p. 62.
- Alfatemi, Seyedeh Mahsan Hoseini, et al. "Analysis of virulence genes among Methicillin Resistant Staphylococcus aureus (MRSA) strains."Jundishapur Journal of Microbiology,Vol. 7, No. 6, 2014, p. e10741.
- Wali, Iman, Nadia Ouda, and Eman El-Seidi. "Mupirocin resistance among methicillin resistant Staphylococcus aureus isolates in an Egyptian hospital."Egyptian Journal of Medical Laboratory Sciences,Vol. 20, No. 2, 2011, pp. 81-92.
- Kehrenberg, Corinna, and Stefan Schwarz. "Distribution of florfenicol resistance genes fexA and cfr among chloramphenicol-resistant Staphylococcus isolates."Antimicrobial Agents and Chemotherapy,Vol. 50, No. 4, 2006, pp. 1156-63.
- Mick, Virginie, et al. "Molecular characterization of resistance to Rifampicin in an emerging hospital-associated methicillin-resistant Staphylococcus aureus clone ST228, Spain."BMC Microbiology,Vol. 10, No. 1, 2010, pp. 1-8.
- O’Neill, Alex J., et al. "Antimicrobial activity and mechanisms of resistance to cephalosporin P1, an antibiotic related to fusidic acid."Journal of Antimicrobial Chemotherapy,Vol. 50, No. 6, 2002, pp. 839-48.
- Thati, Venubabu, Channappa T. Shivannavar, and Subhaschandra M. Gaddad. "Vancomycin resistance among methicillin resistant Staphylococcus aureus isolates from intensive care units of tertiary care hospitals in Hyderabad."The Indian Journal of Medical Research,Vol. 134, No. 5, 2011, pp. 704-08.
- Akpaka, Patrick E., Rashida Roberts, and Stefan Monecke. "Molecular characterization of antimicrobial resistance genes against Staphylococcus aureus isolates from Trinidad and Tobago."Journal of Infection and Public Health,Vol. 10, No. 3, 2017, pp. 316-23.
- San Sit, Pik, et al. "Prevalence of Methicillin-Resistant Staphylococcus aureus (MRSA) infection and the molecular characteristics of MRSA bacteraemia over a two-year period in a tertiary teaching hospital in Malaysia."BMC Infectious Diseases,Vol. 17, No. 1, 2017, pp. 1-14.
- Zainol Abidin, N. Z. B., et al. "MRSA infection in general surgical wards in a Malaysian tertiary hospital: A retrospective study."Annals of Clinical Surgery, Vol. 1, No. 2, 2020, p. 1008.
- Ramazoni, Muhammadji, M. L. Siregar, and K. F. Jamil. "Vancomycin-Resistant Staphylococcus aureus (VRSA) in hepatic cirrhosis patient: A case report."IOP Conference Series: Earth and Environmental Science, Vol. 125, No. 1, 2018, pp.1-3.
- Waitayangkoon, Palapun, et al. "Hospital epidemiology and antimicrobial susceptibility of isolated methicillin-resistant Staphylococcus aureus: A one-year retrospective study at a tertiary care center in Thailand."Pathogens and Global Health,Vol. 114, No. 4, 2020, pp. 212-17.
- He, Wenqiang, et al. "Population structure and characterisation of Staphylococcus aureus from bacteraemia at multiple hospitals in China: Association between antimicrobial resistance, toxin genes and genotypes."International Journal of Antimicrobial Agents,Vol. 42, No. 3, 2013, pp. 211-19.
- VasanthaKumari, Neela, et al. "Prevalence of Panton-Valentine leukocidin genes among carriage and invasive Staphylococcus aureus isolates in Malaysia."International Journal of Infectious Diseases,Vol. 13, No. 3, 2009, pp. e131-32.
- Bariman, Mohammad Hanif, Mohammed Imad A. Mustafa Mahmoud, and Hairul Aini binti Hamzah. "Phenotypic and genotypic characterization, and detection of PVL encoding gene in methicillin resistant Staphylococcus Aureus strains isolated from patients admitted to a tertiary hospital in Kuantan, Malaysia."IIUM Medical Journal Malaysia,Vol. 18, No. 2, 2019, pp. 87-94.
- Samsudin, Syahirah, et al. "Prevalence of Panton-Valentine Leukocidin Staphylococcus aureus among patients and healthcare workers in Malaysian Public Hospital."International Journal of Engineering and Technology,Vol. 7, No. 4.42, 2018, pp. 194-96.
- Shallcross, Laura J., et al. "The role of the Panton-Valentine leucocidin toxin in staphylococcal disease: A systematic review and meta-analysis."The Lancet Infectious Diseases,Vol. 13, No. 1, 2013, pp. 43-54.
- Lim, K. T., et al. "Investigation of toxin genes among methicillin-resistant Staphylococcus aureus strains isolated from a tertiary hospital in Malaysia."Tropical Biomedicine,Vol. 29, No. 2, 2012, pp. 1-8.
- Parsonnet, Jeffrey, et al. "Prevalence of Toxic Shock Syndrome Toxin 1 (TSST-1)-producing strains of Staphylococcus aureus and antibody to TSST-1 among healthy Japanese women."Journal of Clinical Microbiology,Vol. 46, No. 8, 2008, pp. 2731-38.
- Durand, Geraldine, et al. "Detection of new methicillin-resistant Staphylococcus aureus clones containing the toxic shock syndrome toxin 1 gene responsible for hospital-and community-acquired infections in France."Journal of Clinical Microbiology,Vol. 44, No. 3, 2006, pp. 847-53.
- Ghaznavi-Rad, Ehsanollah, et al. "Predominance and emergence of clones of hospital-acquired methicillin-resistant Staphylococcus aureus in Malaysia."Journal of Clinical Microbiology,Vol. 48, No. 3, 2010, pp. 867-72.
- Zarizal, Suhaili, et al. "Nasal colonisation, antimicrobial susceptibility and genotypic pattern of Staphylococcus aureus among agricultural biotechnology students in Besut, Terengganu, east coast of Malaysia."Tropical Medicine and International Health,Vol. 23, No. 8, 2018, pp. 905-13.
- Chen, Hsiao-Jan, et al. "Fusidic acid resistance determinants in Staphylococcus aureus clinical isolates."Antimicrobial Agents and Chemotherapy,Vol. 54, No. 12, 2010, pp. 4985-91.
- Sudagidan, Mert, and Ali Aydin. "Presence of inducible clindamycin resistance phenotype and erm genes in foodborne Staphylococcus aureus isolates."Foodborne Pathogens and Disease,Vol. 10, No. 6, 2013, pp. 555-58.
- Rossato, Adriana Medianeira, Keli Cristine Reiter, and Pedro Alves d’Azevedo. "Coexistence of virulence genes in methicillin-resistant Staphylococcus aureus clinical isolates."Revista da Sociedade Brasileira de Medicina Tropical,Vol. 51, No. 3, 2018, pp. 361-63.
- Yu, Fangyou, et al. "Virulence gene profiling and molecular characterization of hospital-acquired Staphylococcus aureus isolates associated with bloodstream infection."Diagnostic Microbiology and Infectious Disease,Vol. 74, No. 4, 2012, pp. 363-68.