Research - International Journal of Medical Research & Health Sciences ( 2021) Volume 10, Issue 1
Resistance, Phylogenetic Analysis of Pathogenic Escherichia coli in Human Isolate and their Sensitivity with Plant Extracts
Geetika Gupta1*, Ashok Kumar2 and Alok Bhardwaj12Animal Health Division, CIRG, Makhdoom, Farah, Mathura, UP, India
Geetika Gupta, Departments of Biotechnology, Mugla University, Mathura, UP, India, Email: geetikagupta04@gmail.com
Received: 11-Nov-2020 Accepted Date: Jan 22, 2021 ; Published: 29-Jan-2021
Abstract
19 human stool samples were investigated for E. coli pathoyping, phylogenetic analysis, antibiotic resistance pattern, and photosensitivity. 78.94% of samples have the presence of the uspA gene of E. coli. Pathotypes revealed 33.33% Enteropathogenic E. coli (EPEC), 6.66% Enterotoxigenic E. coli (ETEC), but none of the isolates was found positive for EIEC. Phylogenetic markers grouped as (A, B, and D), which were belonged to the D2 group in 40% isolates, 40% isolates belonged to the B group and 20% belongs to the A1 group. Resistances to standard antibiotics/antibacterial were observed with cefixime and cefotaxime 93.33%, 86.66% ceftriaxone and 60% cefepime, 93.33% methicillin, furazolidone (26.66%) and nitrofurantoin 73.33%, sulphadiazine 93.33%,co-trimoxazole 86.66% and trimethoprim 60%, doripenem 46.66%, meropenem and ertapenem 20%, amoxicillin/clavulanic acid, tetracycline and (gatifloxacin) 66.66%, and chloramphenicol 13.33% in human E. coli isolates. Antimicrobial activity with plant extracts (Vitex nigundo, Acacia leucophloea, and Catharanthus roseus) observed at 62.5 mg/ml to 500 mg/ml against the resistant pathogenic isolates. Vitex nigundo were found as the most effective plant extract in ETEC (lt) and EPEC (eae) with a range of zone of inhibition (10-19) mm; and Acacia leucophloea with EPEC (bfpA) exhibited zone of inhibition of (10-17) mm, whereas Catharanthus roseus has poor antimicrobial activity with zone of inhibition of 11 mm or below. This study indicated the higher resistance in E. coli with sulfonamide, tetracycline, and cephalosporin in human kids and the potential of plant extracts for their anti-E. coli activity under in vitro experiments.
Keywords
Enteropathogenic E. coli, Enterotoxigenic E. coli, Phylogenetic groups, ABST, MDR, Zone of Inhibition (ZOI)
Introduction
According to UNICEF, diarrhea is a major cause of child death resulting approximately in 8 percent of all deaths occurs in below age 5 children worldwide in 2017. In sub-Saharan Africa and South Asia, most of the deaths occur in less than 2 years living children from diarrhea [1]. A study reported that childhood diarrheal disease nearly 1.7 billion cases per year globally [2].
Escherichia coli are found in the intestine as normal microflora in humans but sometimes cause disease. Pathogenic strains of E. coli are classified into two groups-intestinal pathogenic E. coli (DEC) and extra-intestinal pathogenic E. coli (ExPEC). Traveler’s diarrhea is mainly caused by DEC.
E. coli can be distributed into four major phylogenetic groups (A, B1, B2, and D) based on phylogenetic analysis and subdivided into seven subgroups A0, A1, B1, B2, B3, D1, and D2. The detection of three primers chu A (the outermembrane hemin receptor gene), yjaA (the gene encodes an uncharacterized protein) and TSPE4.C2 (an anonymous DNA fragment) can be done by using a rapid multiplex method for Polymerase Chain Reaction (PCR) based method [3,4]. A study reported that extraintestinal pathogenic strains of groups B2 and D have virulence factors more commonly, while the majority of A and B1 strains are non-pathogenic and isolated from non-diarrheic fecal samples [5,6].
The most common gram-negative pathogen in humans, Escherichia coli is of particular concern because shows multidrug resistance, and their resistance genes are easily transferable to other strains. Such resistance can spread through fecal contamination of surface water, wetlands, river water, and even drinking water [7-11]. An important mechanism in the development of multidrug-resistant isolates is deprivation and obtains of resistance genes by mobile genetic elements [12]. DEC isolates showed a high prevalence of antibiotic resistance may be due to the acquisition of the resistance genes from intestinal microbiota as reservoirs for carrying of these genes [13]. More than 70 years ago the discovery of antibiotics initiated a period of implementation and drug innovation in animal and human health and agriculture. In all cases, these discoveries were tempered by the appearance of resistant microorganisms. The fact that collections of microorganisms that predate the antibiotic era are highly susceptible to antibiotics reinforced history has been interpreted to mean that antibiotic resistance in pathogenic bacteria is a modern circumstance [14]. An increase in antimicrobial resistance has reached proportions of worldwide magnitude and create a warning to the effective treatment of various infectious diseases [15,16].
Natural products and traditional medicines are of great importance. Traditional medicine is skills and practices, beliefs, the total of knowledge, and experiences indigenous to different cultures that are used to maintain health, as well as to prevent, diagnose, treat or improve mental and physical illnesses. It can treat various infectious and chronic conditions. In some Asian and African countries, the primary health care of 80% population depends on traditional medicine. Alternative or complementary medicine has been used in 70% to 80% of the population in many developed countries [17].
The present study aimed to isolate E. coli from human children of Makhdoom village Mathura, India and to establish correlations among phylotypic groups, resistance, and virulence factors genes profiles. The primary objectives were to (1) measure prevalence of human fecal contamination; (2) measure prevalence of pathogenic E. coli in human young children; (3) examine the effect of antibiotics or group of antibiotics on bacteria in young children, and (4) To study the in vitro antimicrobial activity of different plant extracts against multidrug-resistant human pathogens by disc diffusion method.
Materials and Methods
Collection of Sample
At the time of collection of samples consent letters were signed by parents or guardians as permission to collect sample collection. The fresh fecal material can be put in a container with a stopper or sample container. A total of 19 diarrheic stool samples were collected randomly from human kids (0-3) year at Makhdoom villages, Mathura district.
Transportation
Samples were carried in the icebox to the laboratory after proper labeling.
Storage of Samples
Samples for E. coli isolation were treated immediately after collection normally within 2-3 hours, when not processed immediately in that situation samples were stored at low temperatures for the survival of the bacteria. For future use of samples or repeatedly thawing of samples, pure samples were kept for a long period with glycerol broth at lower temperatures -70°C.
Identification and Confirmation of E. coli
The bacteriological isolation from all samples was done as per the method [18]. Confirmation of E. coli isolates was performed by characteristic biochemical, cultural, and molecular methods (PCR) including characteristics biochemical (IMViC) Catalase test and Oxidase test. Stool sample streaked onto enriched solid Brain Heart Agar media (HIMEDIA, India/BBL Difco, USA) then incubated for 17-19 hrs at 37°C. A single colony from the BHI plate was re-streaked on differential media (Mac Conkey agar media) for identification of lactose sugar fermentation activity and again re-streaked on selective media i.e. Eosin methylene agar (HIMEDIA, India/ BBL Difco, USA) media for confirmation of E. coli. Further, a single colony was also transferred to liquid BHI media for isolation and preservation use. Microscopic examinations were performed from a single colony through the Gram’s staining for morphological identification of E. coli. Biochemical properties for the different biochemical tests were performed such as the catalase test, oxidase test (strips from Sigma), 10% H2O2, and IMViC test from the kit (HIMEDIA, India). The molecular method for E. coli confirmation was performed by a polymerase chain reaction to amplify a fragment of the gene encoding for the highly specific E. coli universal stress protein A (uspA gene) [19-21].
DNA Isolation
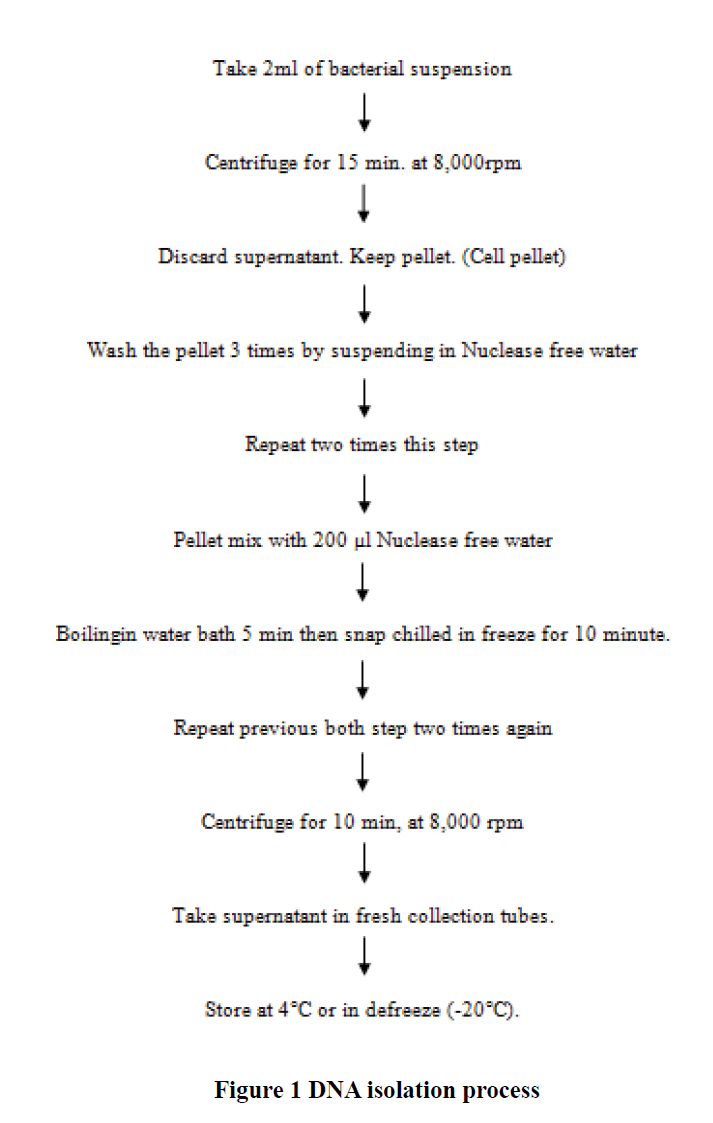
For PCR amplification of the uspA gene and pathotypes of E. coli, the genomic DNA was extracted by the hot-chill method [22]. Briefly, the cell pellet was suspended in 200 μl of nuclease free water and then placed in a boiling water bath for 5 min followed by freezing at -70° for 10 min. Three freeze-thaw cycles were performed, and then centrifugation was done at 8000 g for 10 min. A 50 μl of the supernatant was transferred to a sterile tube and stored at -20°C till further use (Figure 1).
Molecular Confirmation of E. coli and their Pathotypes
The primers (for confirmation) were used in the study as Mishra et al., (Table 1) [21]. PCR reactions were performed in a 25 μl, including 10 μl Emerald master mix (Takara) 1μl (uspA) of each primer for both forward and reverse, 2 μl template DNA, and remaining amount of nuclease-free water.
| Genes | Pathotype | Primer sequence | Size fragment (bp) | Reference |
|---|---|---|---|---|
| (uspA) | CCGATACGCTGCCAATCAGT | 884 | PCR (58°C) Mishra et al., [21] | |
| ACGCAGACCGTAGGCCAGAT | ||||
| ( bfpA) | EPEC | As Catalina et al., [23] | 324 | Multiplex PCR (50°C) Catalina et al., [23] |
| (eae) | EPEC | 384 | ||
| (LT) | ETEC | 450 | ||
| (ST) | ETEC | 190 | ||
| (ial) | EIEC | 650 | ||
|
usp: Universal stress protein; bfp: Bundle forming protein; eae: Intimin; LT: Labile Toxin; ST: Stable Toxin; ial: invasion associate locus; EPEC: Enteropathogenic E .coli; ETEC: Enterotoxigenic E. Coli; EIEC: Enteroinvasive E.coli |
||||
Table 1 The primers used in the study for molecular and pathotypes confirmation of E. coli
The primers (pathotypes) were used in the study as Catalina et al., (Table 1), purchased from Eurofins, USA (Table 2) [23]. Multiplex PCR reactions were performed in a 25 μl, including 10 μl Emerald master mix (Takara) with 0.25 (bfpA) μl, 0.50 (lt) μl, and 0.38 (eae) μl of each primer for both forward and reverse, 2 μl template DNA and remaining amount of nuclease-free water.
| Genes | Primer sequence | Size fragment (bp) | Reference |
|---|---|---|---|
| Chu | As Clermont et al., [3] | 279 | Clermont et al., [3] Multiplex PCR (59°C) |
| Yja | 211 | ||
| TspE4 | 152 |
Table 2 Primer sequences (phylogenic) used in study for Multiplex.
The solution was subjected to thermocycler (peqlab) for defined conditions as per Mishra et al., and Catalina et al., respectively [21,23]. PCR results were analyzed by 1.3% agarose gel electrophoresis and stained by ethidium bromide. Size of bands compared with standard molecular weight marker with 100 bp.
Molecular (Multiplex PCR) Assay for Phylogenic Markers
The Primers (phylogenic) were used in the study as described by Clermont et al., (Table 2) [3].
PCR reactions were performed in a 25 μl, including 10 μl Emerald master mix (Takara) 0.4 μl of each primer for forward and reverse, 2 μl DNA Template and 10.6 μl of nuclease-free water.
The solution was put in a thermocycler (peqlab) for defined conditions at 94°C (4 min,1 cycle); 30 cycles of 94°C (5 sec.), 59°C (10 sec.), and 72°C (30 sec); and a final extension step at 72°C (5 min,1 cycle). Amplified PCR products were analyzed as previously.
Antibiotic Susceptibility Testing
Antimicrobial susceptibility was performed by the Kirby Bauer disk diffusion method on Muller Hinton agar [24]. The antimicrobial agents were tested belonging to groups of cephalosporin, nitrofurans, tetracycline, penicillin, chloramphenicol, penicillin derivative, fluoroquinolones, sulphonamide, and carbapenem group. Antimicrobial susceptibility tests were proceeding on bacterial isolates from stool using antimicrobial discs, according to Clinical and Laboratory Standards Institute (CLSI) guidelines on Muller Hinton agar. The prepared media was inoculated with bacterial suspension equal to 0.5 of McFarland turbidity. The industrially available antimicrobial discs having the following antimicrobials group: cephalosporin, nitrofurans, tetracycline, penicillin, chloramphenicol, penicillin derivative, fluoroquinolones, sulphonamide, and carbapenem group (HIMEDIA, India and Difco, USA) were aseptically placed on the surfaces of the sensitivity agar plates with a sterile forceps and allowed to stay for 30 minutes then incubated at 37°C for (18-24) h. According to CLSI, zones of inhibition after incubation were observed, measured, and interpreted. For each antimicrobial agent used, results were reported as sensitive, intermediate, or resistant. Multidrug-resistant isolates were defined as isolates that demonstrated resistance to ≥3 of the following antimicrobial agent categories as (1) penicillin (AMC-30) (2) nitrofurans (Fr and NIT) (3) cephalosporin (CTX-30, CRO-30 and CTR-30, CFM-5, and FEP-30) (4) sulphonamides (Sz, COT and SXT) (5) tetracycline (TE-30) (6) carbapenems (ETP-10, MRP-10 and DOR-10) (7) chloramphenicol (C-30) (8) penicillin derivative (MET-5), and (9) fluoroquinolones (GAT-5).
Selection and collection of plant material: Three plants were selected for the study. Vitex nigundo L. belongs to the family Verbenaceae. It is a woody, aromatic shrub, commonly known as five-leaved chaste tree in English. Vitex nigundo has been traditionally used in ayurvedic medicine to treat several disorders such as catarrh, headache, neck gland sores, tubercular neck swellings, sinusitis, sexual debilities, nervous debility, liver complaints, inflammation of uterus, fever, diarrhea, etc [25].
A. leucophloea is a medicinal plant employed in the Indian indigenous system of medicine [26,27]. Traditionally; parts of those plants are used against diarrhea and other different problems. The leaves are believed to possess antimicrobial principles [28].
The plant Vinca rosea Linn (periwinkle) is an apocynaceous, ever-blooming, subshrub, or pubescent herb which is a source of many alkaloids. Catharanthus roseus is an erected procumbent herb or under the shrub containing latex, rich in their pharmacological actions that include antibacterial, anticancer, antioxidant, antifungal, and antiviral activates [29]. Externally, it is used in case of mouth ulcers, nosebleeds, sore throats, and bleeding gums. It is also used internally for diarrhea, gastritis, and enteritis, cystitis, raised blood sugar levels, hypertension, and loss of memory [30].
Leaves of the above-selected plants were collected from the Institute medicine garden at Mathura UP, India. The plant material was cleaned and dried at room temperature. After drying, leaves were ground in an electronic grinder for fine & coarse powder. Powders were kept in airtight containers for future use.
Preparation of extract: 80 g of crushed material was extricated in 800-milliliter methanol by soxhlet apparatus at (40-50)°C, followed by vacuum evaporation for the elimination of methanol. After drying crude extract was collected and preserved in an airtight storage container for further use. The crude extract was used at the highest concentrations of 500 mg/ml which was further serially, diluted to the concentrations of 62.5 mg/ml, 125 mg/ml, and 250 mg/ml.
Herbal sensitivity testing/antimicrobial assay of plant extracts: Antimicrobial assay of extracts of different plants was performed by the Disc diffusion method in Mueller Hinton Agar (MHA) plates with the help of sterile discs (HIMEDIA, India). The test bacteria were inoculated in BHI broth and incubated overnight at 37°C to justify the turbidity to 0.5 McFarland standards giving a final inoculum of 1.5 × 108 CFU/ml. MHA plate was lawn cultured. Plant extracts of 500 mg/ml concentration were prepared in methanol (MeOH) and prepare their dilutions (250, 125, 62.5) mg/ml respectively. MHA plate inoculated with standardized microbial culture broth and sterile disc (6 mm) were filled with prepared plant concentrations. It was permitted to absorb at room temperature for about 30 minutes after that discs filled with plant extracts were applied on the inoculated MHA plate inoculated with microbial culture and incubated for (18-24) hours at 37°C. Plates were observed after incubation for the formation of a zone around the disc which corresponds to the antimicrobial activity of tested compounds. The Zone of Inhibition (ZOI) was observed and measured in mm
Discussion
Microbiological Determination
E. coli has been reported as the important causative agent of diarrhea in humans as well as in animals. Isolation and identification E. coli were done based on cultural, morphological, and biochemical characteristics [18]. The organisms were Gram-negative short rods (coccobacilli), catalase-positive, oxidase negative, and IMViC reactions as (+, +, - and-). Lysine utilization, ornithine utilization, nitrate reduction, glucose, lactose, and arabinose were positive. Urease, phenylalanine deamination, H2S production, adonitol, and sorbitol were negative. In our results, E. coli demonstrated 78.94% with uspA gene in human kid samples, while in a study occurrence of E. coli in stool samples of healthy humans reported 92.68% [33].
Pathogenic Group Determination
Results showed that 33.33% (5 samples) positive isolates belonged to EPEC and one sample to ETEC (6.66%). In which two positive isolates belonged to EPEC (eae) and three samples to EPEC (bfpA), whereas ETEC (st) and EIEC were not detected in human kids in this study (Table 4). In another such study reported that EPEC (50.0%) was the most frequent pathotypes followed by EAEC (20.4%), ETEC (14.8%), EIEC (3.7%), and STEC (3.7%) [34].
| No. of isolates | EPEC (bfpA) | EPEC (eae) | ETEC (LT) | ETEC (st) | EIEC (ial) |
|---|---|---|---|---|---|
| 15 | 2 (13.33%) | 3 (20%) | 1 (06.66%) | 0 | 0 |
Table 4 Percent occurrence of E. coli pathotypes
| No. of isolates | D2 | B1 | B2 | A1 |
|---|---|---|---|---|
| 15 | 6 (40%) | 4 (26.66%) | 2 (13.33%) | 3 (20%) |
Table 5 % Occurrence of E. coli phylogenic groups.
Phylogenetic Group Determination
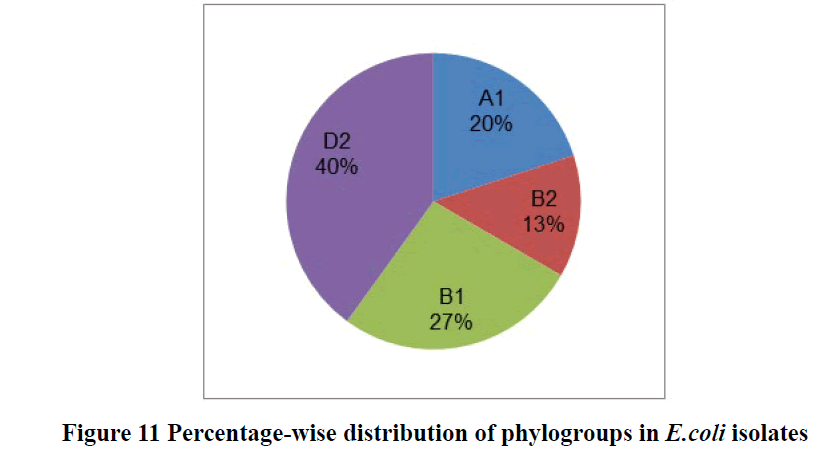
The phylogenetic marker represented as chu A for extraintestinal E. coli isolates, whereas yjaA and TspE4.C2 for intestinal E. coli isolates. Results showed that most of the E.coli isolates were belonged to (40%) D2 phylogenic group followed by 26.66% B1 group, 13.33% B2, and 20% A1 respectively. Phylogenic group showed as D2 (chu+ and TspE4. C2+), B1 (TspE4.C2+), B2 (chu+ and yja+) and A1 (yja+) (Table 5 and Figure 11). Our study results revealed that most of the isolates belonged to extraintestinal E. coli (53.33%) and (46.66%) to commensal pathogenic or non-pathogenic intestinal E. coli in human kids.
In other similar studies had demonstrated that B2 are most common extraintestinal pathogenic strains in isolates and A and B1 phylogenetic group isolates were strictly belonging to commensal strains from the intestinal microbiota while Ghazal Naderi et al., reported 13.2% of DEC isolates from A and B1 groups commented that these groups are also eligible of causing diarrheal diseases [35-37]. The results also showed that group B2, despite being uncommon among commensal isolates, can be present in intestinal flora (18% of our commensal isolates), suggesting that they may act as a reservoir for bacteria that can cause diarrheal diseases. This study has reported different findings, where the predominant phylogenetic group in UPEC isolates was D2 group (40%) followed by B1 group (26.66%), B2 (13.33%), and lastly A1 (20%). our observations were more similar to these results.
In another study, commented that commensal E. coli isolates showed the occurrence of phylogroups such as B2 group 9 (30%), D group 12 (40%), and A groups 9 (30%), respectively but none of the isolates tested clustered in B1 group [33]. These results have approximate similarities with our results.
Antibiotic Resistance Pattern
In another study, the high incidence of amoxicillin/clavulanic acid (58.1%) resistance was reported in stools from young children as this study also showed 66.66% resistance to amoxicillin/clavulanic acid antibiotic [37]. However, 19% of the isolates showed resistance to clavulanic acid combined with amoxicillin (amoxicillin-clavulanic acid) in environmental samples from hospitals and homes so here our results mismatch with the study [38]. According to Ghanem and Haddadin, the majority of the isolates (71.4%) exhibited resistance to trimethoprim-sulfamethoxazole, to a cephalosporin (second and third generation) was ranged from 57% to 66.7% and considerable and no resistance was detected against imipenem, while in our results only 60% resistance occurred to trimethoprim-sulfamethoxazole and (60-94)% to cephalosporins [38].
Our results also showed D2 group occurred predominantly and all pathotypes are multi-drug resistant to more than three antibiotics and all groups showed resistance tocefotaxime, ceftriaxone, tetracycline, and cefixime (Table 6).
| Pathogenic isolates | Phylogenic group | Resistant to antibiotics |
|---|---|---|
| EPEC | D2 | COT,CTX,SZ,CFM,NIT,TE,AMC,C,CRO,FEP,GAT,MET |
| EPEC | D2 | CTX,CFM,TE,AMC,CRO,GAT,MET,CAZ |
| ETEC (lt) | B1 | MRP, ETP, COT, CTX, CTR, SZ, CFM, NIT, TE, AMC, SXT, CRO |
| EPEC (bfpA) | A1 | COT, CTX,CTR, Sz, Fr, CFM, NIT, TE, SXT, CRO, FEP, MET, CAZ |
| Ertapenem (ETP-10); Meropenem (MRP-10); Cefotaxime (CTX-30); Ceftriaxone (CRO-30 and CTR-30); Cefixime (CFM-5); Cefepime (FEP-30); Tetracycline (TE-30); Sulphdiazene (Sz); Co-trimoxazole (COT); Sulphamethoxazole with trimethoprim (SXT); Nitrofurantoin (NIT); Furazolidone (Fr); Amoxicillin/Clavulanic Acid (AMC-30); Chloramphenicol (C-30); Gatifloxacin (GAT-5) and Methicillin (MET-5) | ||
Table 6 Occurrence of multidrug-resistant E. coli phylogenic groups in pathogenic isolates.
Antimicrobial Sensitivity of Plant Extracts
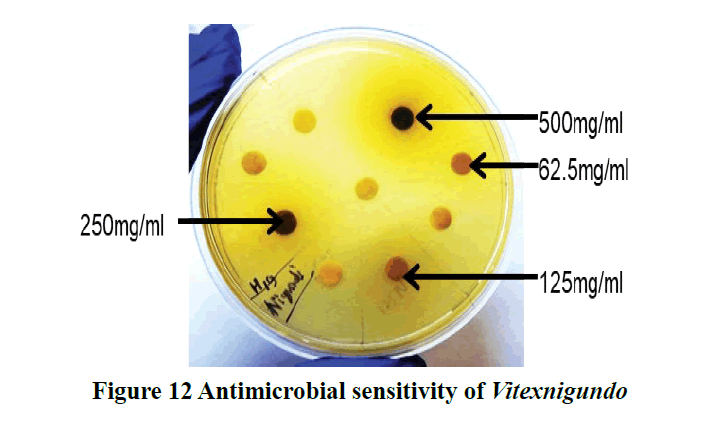
In a study, promising antibacterial activity of nigundo has been reported in ethanolic and methanolic leaves extract against both gram-positive and gram-negative bacteria (E. coli) [39]. In another study, the antibacterial effect of nigundo against E. coli have been found at all concentration and the highest ZOI was 12 mm (100 mg/ml) while in our results Vitex nigundo showed the highest Zone of Inhibition (ZOI) 19 mm (500 mg/ml) (Figure 12) [40]. Here our results differ in the effectiveness of plant extract at concentration. Another study has been reported that the zones of inhibition produced by the methanol extract fraction were found to be (7-16) mm at a 200 μg/disc concentration [41].
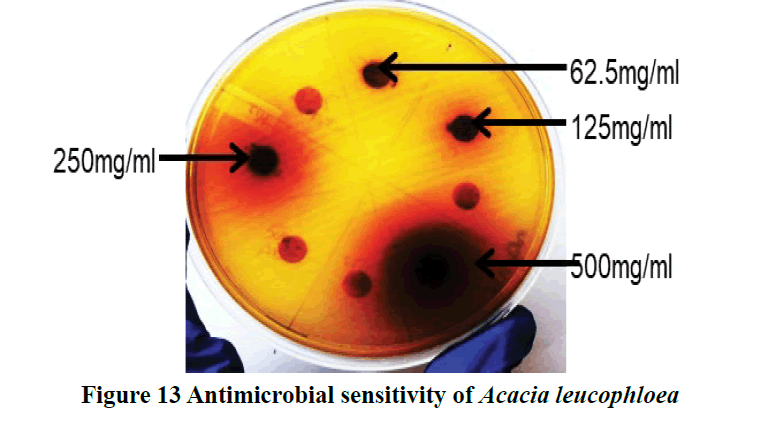
Gilani et al., also reported a methanolic extract of Acacia leucophloea exhibited dose-dependent (100-500) mg/ml protective effect against castor oil-induced diarrhea in mice as shown by a decrease in the total number of feces [42]. The significant anti-diarrheal activity showed by methanolic extract of Acacia leucophloea. This was expected as anticholinergic drugs and Ca2+ antagonists possess anti-diarrheal activity [43,44]. While in our findings Acacia leucophloea showed the highest Zone of Inhibition (ZOI) 16 mm (500 mg/ml) (Figure 13). Here are results similar in the effectiveness of plant extract at concentration.
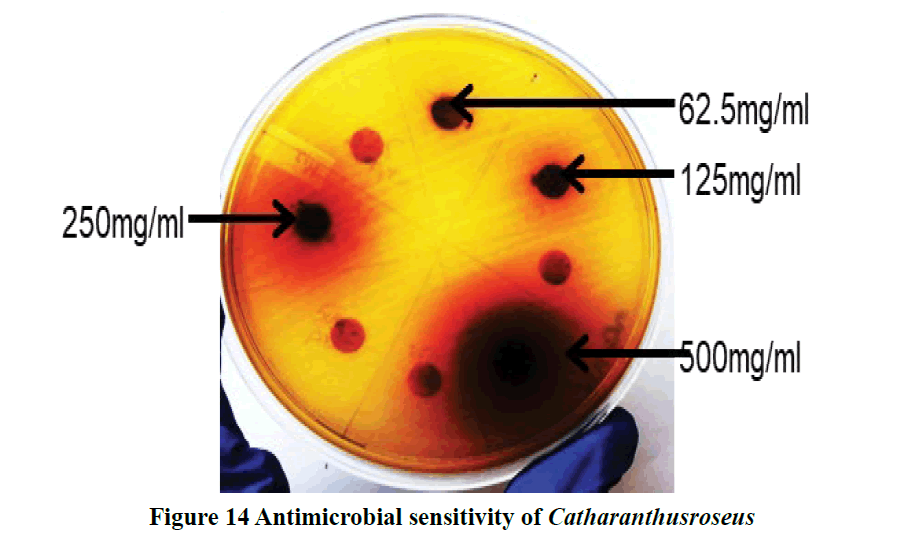
The anti-diarrheal effect of ethanolic extracts C. roseus showed the dose-dependent inhibition of the castor oil-induced diarrhea (Figure 14) [45].
Conclusion
E. coli is an environmental pathogen affecting human kids, mainly causing diarrhea. EPEC pathotype of E. coli occurred most frequently in human kids, belonged to extraintestinal pathotype and D2 phylogenic group. E. coli isolates from human kids showed resistance to different classes of commonly used antibiotics such as cephalosporin, sulfonamide, and penicillin derivative, and nitrofurans, which warrant the highest risk and need all-time attention. Alternative medicine by using medicinal plants such as Acacia and Vitex plant leaves can be a potential approach.
Declarations
Funding
This study was funded by the Indian council of agriculture research, New Delhi.
Conflicts of Interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical requirements declaration
Stool samples used in studies involving children were in accordance with the approval of his parents at which the studies were conducted.
Informed consent
Informed consent was obtained from all individual participant parents included in the study.
Author’s contributions
Compilation of data, paper writing work with figures and tables finalized by Geetika Gupta and Dr. Ashok Kumar checked and finalized the research paper with some correction. Dr. Alok Bhardwaj checked plagiarism in the paper.
References
- UNICEF, WHO. "Diarrhoea remains a leading killer of young children, despite the availability of a simple treatment solution." UNICEF data: monitoring the situation of children and women, 2016.
- World Health Organization. Diarrhoeal disease. Geneva: World Health Organization, 2017.
- Clermont, Olivier, Stéphane Bonacorsi, and Edouard Bingen. "Rapid and simple determination of theEscherichia coli phylogenetic group." Applied and Environmental Microbiology, Vol. 66, No. 10, 2000, pp. 4555-8.
- Kalantar, Enayatolah, et al. "Antibiotic resistance patterns of STEC and ETEC strains: a study on frozen foods of animal origin and children with acute diarrhea." Journal of Microbiology and Infectious Diseases, Vol. 3, No. 1, 2013, pp. 31-5.
- Duriez, Patrick, et al. "Commensal Escherichia coli isolates are phylogenetically distributed among geographically distinct human populations." Microbiology, Vol. 147, No. 6, 2001, pp. 1671-6.
- Johnson, James R., et al. "Phylogenetic distribution of extraintestinal virulence-associated traits in Escherichia coli." The Journal of Infectious Diseases, Vol. 183, No. 1, 2001, pp. 78-88.
- Vivant, Anne-Laure, et al. "Free water surface constructed wetlands limit the dissemination of extended-spectrum beta-lactamase producing Escherichia coli in the natural environment." Water Research, Vol. 104, 2016, pp. 178-88.
- Rodriguez-Mozaz, Sara, et al. "Occurrence of antibiotics and antibiotic resistance genes in hospital and urban wastewaters and their impact on the receiving river." Water Research, Vol. 69, 2015, pp. 234-42.
- Li, Xunde, et al. "Antibiotic-resistant E. coli in surface water and groundwater in dairy operations in Northern California." Environmental Monitoring and Assessment, Vol. 186, No. 2, 2014, pp. 1253-60.
- Coleman, Brenda L., et al. "Contamination of Canadian private drinking water sources with antimicrobial resistant Escherichia coli." Water Research, Vol. 47, No. 9, 2013, pp. 3026-36.
- Baquero, Fernando, José-Luis Martínez, and Rafael Cantón. "Antibiotics and antibiotic resistance in water environments." Current Opinion in Biotechnology, Vol. 19, No. 3, 2008, pp. 260-5.
- Wozniak, Rachel AF, and Matthew K. Waldor. "Integrative and conjugative elements: mosaic mobile genetic elements enabling dynamic lateral gene flow." Nature Reviews Microbiology, Vol. 8, No. 8, 2010, pp. 552-63.
- Ravi, Anuradha, et al. "Integrons in the intestinal microbiota as reservoirs for transmission of antibiotic resistance genes." Pathogens, Vol. 3, No. 2, 2014, pp. 238-48.
- D’Costa, Vanessa M., et al. "Antibiotic resistance is ancient." Nature, Vol. 477, No. 7365, 2011, pp. 457-61.
- Centres for Disease Control and Prevention (US). Antibiotic resistance threats in the United States, 2013. Centres for Disease Control and Prevention, US Department of Health and Human Services, 2013.
- World Health Organization. "Antimicrobial resistance: global report on surveillance." World Health Organization, 2014.
- Rupasinghe, L. "Traditional Medicine in Asia: World Health Organization." New Delhi: Regional Office for South-East Asia, 2001.
- Cowan, Samuel Tertius, and Kenneth John Steel. "Manual for the identification of medical bacteria." Manual for the Identification of Medical Bacteria, 1965.
- Osek, Jacek. "Multiplex polymerase chain reaction assay for identification of enterotoxigenic Escherichia coli strains." Journal of Veterinary Diagnostic Investigation, Vol. 13, No. 4, 2001, pp. 308-11.
- Rajput, S. K., et al. "Study of the characterization of E. coli isolates in goat kids." Indian Research Journal of Genetics and Biotechnology, Vol. 6, No. 1, 2014, pp. 324-9.
- Mishra, Anil K., et al. "UspA gene based characterization of Escherichia coli strains isolated from different disease conditions in goats." Journal of Animal Research, Vol. 7, No. 6, 2017, pp. 1123-8.
- Yang, Jin-Long, et al. "A simple and rapid method for extracting bacterial DNA from intestinal microflora for ERIC-PCR detection." World Journal of Gastroenterology: WJG, Vol. 14, No. 18, 2008, pp. 2872.
- López-Saucedo, Catalina, et al. "Single multiplex polymerase chain reaction to detect diverse loci associated with diarrheagenic Escherichia coli." Emerging Infectious Diseases, Vol. 9, No. 1, 2003, pp. 127.
- Bauer, A. W. "Antibiotic susceptibility testing by a standardized single disc method." American Journal of Clinical Pathology, Vol. 45, 1966, pp. 149-58.
- Ladda, P. L., and C. S. Magdum. "Vitex negundo Linn.: Ethnobotany, phytochemistry and pharmacology-A review." International Journal of Advances in Pharmacy, Biology and Chemistry, Vol. 1, No. 1, 2012, pp. 111-20.
- Chopra, Ram Nath, Sham L. Nayar, and Ishwar C. Chopra. "Glossary of Indian medicinal plants." Vol. 1. New Delhi: Council of Scientific & Industrial Research, 1956.
- Kirtikar, K. R. B. B., and B. D. Basu. "Indian medicinal plants." Indian Medicinal Plants, 1935.
- Khare, C. P. "Indian Medicinal Plants: An Illustrated Dictionary: Springer." 2007.
- Gajalakshmi, S., S. Vijayalakshmi, and Rajeswari V. Devi. "Pharmacological activities of Catharanthus roseus: a perspective review." International Journal of Pharma and Bio Sciences, Vol. 4, No. 2, 2013, pp. 431-9.
- Balaji, Dr Hemamalini. "Versatile. Therapeutic effects of Vinca rosea Linn." International Journal of Pharmaceutical Science and Health Care, Vol. 1, No. 4, 2014, pp. 59-76.
- Abdul-Razzaq, Mohammad S., and Lamees A. Abdul-Lateef. "Molecular phylogeny of Escherichia coli isolated from clinical samples in Hilla, Iraq." African Journal of Biotechnology, Vol. 10, No. 70, 2011, pp. 15783-7.
- Khairy, Rasha M., et al. "Phylogenic classification and virulence genes profiles of uropathogenic E. coli and diarrhegenic E. coli strains isolated from community acquired infections." PLoS One, Vol. 14, No. 9, 2019, pp. e0222441.
- Mirzarazi, Mahsa, et al. "Occurrence of genes encoding enterotoxins in uropathogenic Escherichia coli isolates." Brazilian Journal of Microbiology, Vol. 46, No. 1, 2015, pp. 155-9.
- Zhou, Yu, et al. "Characteristics of diarrheagenic Escherichia coli among children under 5 years of age with acute diarrhea: a hospital based study." BMC Infectious Diseases, Vol. 18, No. 1, 2018, pp. 63.
- Li, Bin, et al. "Phylogenetic groups and pathogenicity island markers in fecal Escherichia coli isolates from asymptomatic humans in China." Applied and Environmental Microbiology, Vol. 76, No. 19, 2010, pp. 6698-700.
- Koga, Vanessa L., et al. "Molecular screening of virulence genes in extraintestinal pathogenic Escherichia coli isolated from human blood culture in Brazil." BioMed Research International, Vol. 2014, 2014.
- Naderi, Ghazal, et al. "Distribution of pathogenicity island (PAI) markers and phylogenetic groups in diarrheagenic and commensal Escherichia coli from young children." Gastroenterology and Hepatology from Bed to Bench, Vol. 9, No. 4, 2016, pp. 316.
- Ghanem, Bothyna, and Randa Nayef Haddadin. "Multiple drug resistance and biocide resistance in Escherichia coli environmental isolates from hospital and household settings." Antimicrobial Resistance and Infection Control, Vol. 7, No. 1, 2018, pp. 47.
- Thatoi, H. N., and S. K. Dutta. "Antibacterial activity and phytochemical screening of leaf and bark extracts of Vitex negundo l. from similipal biosphere reserve, Orissa." Journal of Medicinal Plants Research, Vol. 3, No. 4, 2009, pp. 294-300.
- Deogade, Meena Shamrao, et al. "Antimicrobial Activity of Vitex Negundo Linn. (Nirgundi) Leaves Extract." Journal of Research in Traditional Medicine, Vol. 2, No. 4, 2016, pp. 99-102.
- Chowdhury, N. Y., W. Islam, and M. Khalequzzaman. "Insecticidal activities of the leaves of nishinda (Vitex negundo L. Verbinaceae) against Tribolium castaneum Hbst." Pakistan Entomologist, Vol. 31, No. 1, 2009, pp. 25-31.
- Gilani, Anwarul Hassan, et al. "Mechanisms underlying the antispasmodic and bronchodilatory properties of Terminalia bellerica fruit." Journal of Ethnopharmacology, Vol. 116, No. 3, 2008, pp. 528-38.
- Reynolds, I. J., ROBERT J. Gould, and Solomon H. Snyder. "Loperamide: blockade of calcium channels as a mechanism for antidiarrheal effects." Journal of Pharmacology and Experimental Therapeutics, Vol. 231, No. 3, 1984, pp. 628-32.
- Rang, H. P., et al. "Pharmacology, 6. ed." Churchill Livingstone/Elsevier, 2007, pp. 692-7.
- Rajput, Mithun Singh, et al. "Evaluation of antidiarrheal activity of aerial parts of Vinca major in experimental animals." Middle-East Journal of Scientific Research, Vol. 7, No. 5, 2011, pp. 784-8.