Research - International Journal of Medical Research & Health Sciences ( 2021) Volume 10, Issue 4
Molecular Characterization of Colistin Resistant Klebsiella Pneumoniae ST11 Responsible for Nosocomial Outbreak in Neonatal Intensive Care Unit, India
Santosh K Singh*Santosh K Singh, Department of Biotechnology, ARKA Jain University, Jamshedpur, Jharkhand, India, Email: thatoinou@gmail.com
Received: 25-Mar-2021 Accepted Date: Apr 20, 2021 ; Published: 27-Apr-2021, DOI: 0
Abstract
The dramatic increase of resistance in Klebsiella pneumoniae (K. pneumoniae) due to the misuse of antibiotics poses one of the greatest health threats and is a global health concern. Colistin, a last resort antibiotic is used extensively to treat carbapenem-resistant Klebsiella pneumoniae infections. Present research carried out to analyze the molecular mechanism, clonal types, and outcomes of the infections caused by colistin-resistant K. pneumoniae in neonates during an outbreak in neonate intensive care unit. Twenty-eight cases of colistin-resistant, carbapenem-resistant K. pneumoniae were identified between March and April 2016. Isolates were genotyped using multi-locus sequence typing and the molecular mechanism of colistin resistance was ascertained. All the colistin-resistant K. pneumoniae isolated from neonates during outbreak have insertional inactivation by ISL3 family transposes in the mgrB gene and were clonally related belong to ST11. PCR screening confirmed the presence of the blaOXA-48 and blaSHV-34 genes. The observed mortality was 35.7% in two-month periods. The present baseline report of colistin-resistant K. pneumoniae ST11 outbreak suggested the emergence of clones with this phenotype that required paramount importance for future health monitoring and assessment.
Keywords
Antibiotics, β-lactamase, Colistin-resistance, Klebsiella pneumoniae, plasmid
Introduction
Klebsiella pneumoniae has emerged as a superbug and poses as one of the world’s greatest health threats [1]. It is the causative agent of a variety of diseases, commonly including urinary tract, soft tissue infections, bacteremia, and pneumonia; however, community-acquired invasive infections have also emerged as of late in the form of a pyogenic liver abscess, brain abscess, meningitis, and ankylosing spondylitis [2,3]. Eradication of this superbug is extremely difficult as it has acquired resistance to almost all available antibiotics and is consequently associated with high morbidity and mortality, thus it has been included among the six multidrug-resistant ESKAPE pathogens (Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter species) [4]. Neonates are more vulnerable to infection due to their immature immune systems. Low birth weight, prematurity, febrile illness in the mother, rupture of membranes, unhygienic conditions, delays in recognition and prolong hospitalization are some recognized risk factors. In India, neonatal septicemia is ported incidence rate of 30 cases per 1000 live births, with K. pneumoniae as the most frequently isolated pathogen in both intramural births (32.5%) and extramural neonates (27%) [5]. The use of cephalosporin and carbapenem leaped forward the fight against infections caused by K. pneumoniae. However, co-production of multiple β-lactamases, especially ESBLs (e.g. blaSHV, blaTEM, and/or blaCTX) and carbapenemase (blaKPC and/or blaOXA), in K. pneumoniae has been reported from many countries, rendering β-lactam antibiotics completely futile [6-8]. The situation is further complicated as Extended-Spectrum β-Lactamase (ESBL)-producing K. pneumoniae isolates exhibited very high (up to 56%) resistance to quinolones [9].
Due to the unavailability of therapeutic alternatives for the treatment of Carbapenem-Resistant K. pneumoniae (CRKP), colistin has been extensively used instead. Colistin Sulfate (CS) is used for oral and topical therapy, whereas Colistin Methanesulfonate Sodium (CMS) is used for parenteral and aerosol therapy [10,11]. Colistin being positively charged, alters the permeability of the bacterial cell membrane by displaying divalent cations, Ca2+and Mg2+, from the phosphate groups of lipid A, thus causing an outflow of cell contents which ultimately results in bacterial death [12]. However, with excessive use, colistin resistance in K. pneumoniae began to increase, which highlights an emerging threat in the treatment of healthcare-related infections [13,14]. PhoP/PhoQ and PmrA/PmrB are the main systems responsible for the modification of the Lipopolysaccharide (LPS) layer. Changes in these regulatory systems cause colistin resistance. Insertional inactivation/mutation of the mgrB gene, which encodes the key regulatory protein of the PhoP/PhoQ system, or the presence of a plasmid-mediated mcr-1 gene are the primary mechanisms driving colistin resistance [15-17].
Between March-July 2016, an outbreak of colistin-resistant K. pneumoniae bacteremia was reported in the neonatal intensive care unit, Jamshedpur, Jharkhand [18]. Keeping the progressive resistance mechanism of K. pneumoniae in mind which is a global health concern, the present research investigation has been carried out to analyze the molecular mechanism of colistin resistance, clonal types, clinical futures, and outcomes of the infections caused by colistinresistant K. pneumoniae isolated from neonates during this outbreak. Our findings will set the stage for future research on this emerging health crisis.
Methods
Bacterial Isolates
Thirty (28 from neonates and 2 from hospital environment) non-duplicate Colistin-Resistant Carbapenem-Resistant K. pneumoniae (COLR-CRKP) isolates from the neonatal intensive care unit of a tertiary care hospital (Jharkhand, India) during an outbreak were included in this study. One ml of blood was drawn from the neonates suspected to have sepsis and inoculated in the BacT/ALERT (bioMerieux, USA) microbial detection system. After the prediction of an outbreak, surveillance swabs from the nursery (air, warmer, suction, IV/PIC line, and oxygen nozzle), gynecology OT (table, light, and instruments), and labor room were taken for culture. Random hand swabs of staff, doctors, and mothers were also taken for culture. Identification and antibiotic susceptibility of isolates were determined by aVITEK® 2 compact system (bioMerieux, USA) using the ID-GNB and AST-N280 cards following the manufacturer’s instructions. The antimicrobial susceptibility testing card included the following antibiotics: Ampicillin (AMP), Cefuroxime (CXM), Ceftriaxone (CRO), Cefepime (FEP), Imipenem (IPM), Meropenem (MEM), Cefoperazone/ Sulbactam (CSS), Piperacillin-Tazobactam (TZP), Amoxicillin-Clavulanic Acid (AMC), Ciprofloxacin (CIP), Trimethoprim- Sulfamethoxazole (SXT), Amikacin (AMK), Gentamicin (GEN), Nitrofurantoin (NIT), Ertapenem (ERT), Co-Trimoxazole (COT) and Colistin (COL). Ertapenem resistance was defined as a MIC of ≥ 8 mg/l and colistin resistance was defined as a MIC of ≥ 4 mg/l [19]. Escherichia coli ATCC 25922 and Klebsiella pneumoniae ATCC 700603 were used as quality control strains for each batch of MIC tests.
Patients and Clinical Epidemiology
Demographic data such as gender, age, clinical diagnosis, location, clinical outcome, and the time of admission, the time of discharge, and the length of hospital stay were extracted from the patient administration system.
Detection of β-lactamase Genes
Phenotypic detection of ESBLs, MBLs, and carbapenemase was carried out using a double disk synergic test, and imipenem EDTA double-disc synergy test, and Modified Hodge Test (MHT) respectively, following the protocol as described by CLSI 2014 manual [20]. The phenotypic detection of AmpC was performed using a ceftazidime-boronic acid combined disc diffusion test as described by Yilmaz, et al. [21]. PCR amplification was carried out in a thermal cycler (Eppendorf, Germany) to detect the genes encoding for blaTEM, blaSHV, blaIMP, blaVIM, blaNDM-1, blaOXA-48, blaKPC, and AmpC according to their respective product size. PCR products were separated in a 1% agarose-TAE gel containing ethidium bromide and visualized in a Chemidoc imager (Bio-Red, USA).
PCR Amplification of mgrB and mcr-1 Genes
PCR amplification of the mgrB gene and mcr-1 gene were carried out to determine the colistin resistance mechanism. The presence of plasmid DNA of all the K. pneumoniae isolates were screened using QIAGEN plasmid mini kit (QIAGEN GmbH, Germany). Amplified PCR products were purified with a QIAquick PCR Purification Kit (QIAGEN) and sequenced with an ABI 3100 Sequencer (Applied Biosystems, Foster City, CA). The nucleotide and deduced protein sequences were analyzed using the NCBI website. The Insertion Sequence (IS) was analyzed using the IS finder website.
Molecular Typing
Strain typing of K. pneumoniae isolates was performed by ERIC-PCR using primer setERIC-2/ERIC-1026 according to Munoz, et al. [22]. Multilocus sequence typing of isolates was performed as described by Diancourt, et al. and analyzed using the MLST website for K. pneumoniae [23]. All the seven house-keeping gene sequences were submitted to get the gene bank accession number.
Results
A total of 28 colistin-resistant K. pneumoniae were isolated between March and April 2016 from the blood culture of neonates admitted to the neonatal intensive care unit (Table 1). Out of 28 neonates, 57% (n=16) were males and 43% (n=12) were females. Besides two more colistin-resistant K. pneumoniae strains (EKpCR1 and EKpCR2) were isolated from the gynecological oxygen nozzle and suction of the Operation Table (OT) of the same hospital. However, all the K. pneumoniae isolates showed uniform antibiogram, pattern, and resistance to the first and second line of cephalosporins, β-lactam/β-lactamase inhibitor combination drugs, carbapenems, aminoglycosides, quinolones, and colistin. The only drugs which were sensitive in all of the isolates were tigecycline and co-trimoxazole, which revealed that all of the isolates belonged to a single clone.
| Isolates ID | Date of isolation of K. pneumoniae | MIC (mg/l) | Gender of patient | A hospital stay of the patient (days) | Outcome of patient | |||
|---|---|---|---|---|---|---|---|---|
| Col | ERT | MER | TGC | |||||
| CKpCR1 | 2-Mar-16 | >16 | >8 | >16 | 2 | F | 33 | Alive |
| CKpCR2 | 26-Mar-16 | >16 | >8 | >16 | 2 | M | 7 | Alive |
| CKpCR3 | 28-Mar-16 | >16 | >8 | >16 | 2 | F | 5 | Dead |
| CKpCR4 | 10-Apr-16 | >16 | >8 | >16 | 2 | M | 7 | Dead |
| CKpCR5 | 10-Apr-16 | >16 | >8 | >16 | 2 | M | 23 | Alive |
| CKpCR6 | 1-Mar-16 | >16 | >8 | >16 | 2 | F | 4 | Alive |
| CKpCR7 | 16-Apr-16 | >16 | >8 | >16 | 2 | F | 26 | Dead |
| CKpCR8 | 5-Mar-16 | >16 | >8 | >16 | 2 | M | 5 | Dead |
| CKpCR9 | 21-Apr-16 | >16 | >8 | >16 | 2 | F | 2 | Alive |
| CKpCR10 | 16-Apr-16 | >16 | >8 | >16 | 2 | F | 29 | Alive |
| CKpCR11 | 14-Mar-16 | >16 | >8 | >16 | 2 | M | 10 | Alive |
| CKpCR12 | 26-Mar-16 | >16 | >8 | >16 | 2 | F | 7 | Dead |
| CKpCR13 | 29-Mar-16 | >16 | >8 | >16 | 2 | M | 11 | Dead |
| CKpCR14 | 16-Apr-16 | >16 | >8 | >16 | 2 | F | 8 | Alive |
| CKpCR15 | 22-Mar-16 | >16 | >8 | >16 | 2 | M | 9 | Alive |
| CKpCR16 | 26-Mar-16 | >16 | >8 | >16 | 2 | M | 7 | Dead |
| CKpCR17 | 6-Apr-16 | >16 | >8 | >16 | 2 | M | 15 | Alive |
| CKpCR18 | 30-Mar-16 | >16 | >8 | >16 | 2 | M | 7 | Alive |
| CKpCR19 | 16-Apr-16 | >16 | >8 | >16 | 2 | M | 18 | Alive |
| CKpCR20 | 13-Mar-16 | >16 | >8 | >16 | 2 | F | 13 | Alive |
| CKpCR21 | 13-Mar-16 | >16 | >8 | >16 | 2 | F | 21 | Alive |
| CKpCR22 | 19-Apr-16 | >16 | >8 | >16 | 2 | M | 6 | Dead |
| CKpCR23 | 12-Apr-16 | >16 | >8 | >16 | 2 | F | 16 | Alive |
| CKpCR24 | 10-Apr-16 | >16 | >8 | >16 | 2 | M | 4 | Alive |
| CKpCR25 | 10-Apr-16 | >16 | >8 | >16 | 2 | M | 10 | Alive |
| CKpCR26 | 26-Mar-16 | >16 | >8 | >16 | 2 | F | 15 | Alive |
| CKpCR27 | 12-Apr-16 | >16 | >8 | >16 | 2 | M | 10 | Dead |
| CKpCR28 | 28-Mar-16 | >16 | >8 | >16 | 2 | M | 7 | Dead |
| #EKpER1 | 4-Apr-16 | >16 | >8 | >16 | 2 | - | - | - |
| #EKpER2 | 4-Apr-16 | >16 | >8 | >16 | 2 | - | - | - |
MIC: Minimum Inhibitory Concentration; Col: Colistin; ERT: Ertapenem; MER: Meropenem, TGI: Tigecyclin
#Colistin resistant and carbapenem-resistant K. pneumoniae from the hospital environment
The isolates were phenotypically negative for ESBL, MBL, and AmpC production, whereas determined to be positive for the production of carbapenemases by a modified Hodge test. PCR screening confirmed the presence of the blaOXA-48 and blaSHV-34 genes.
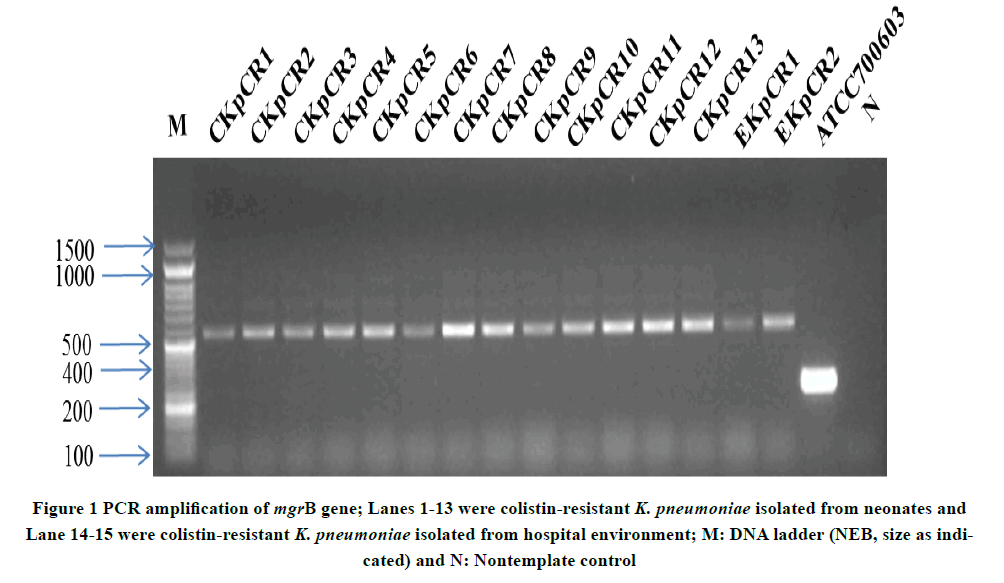
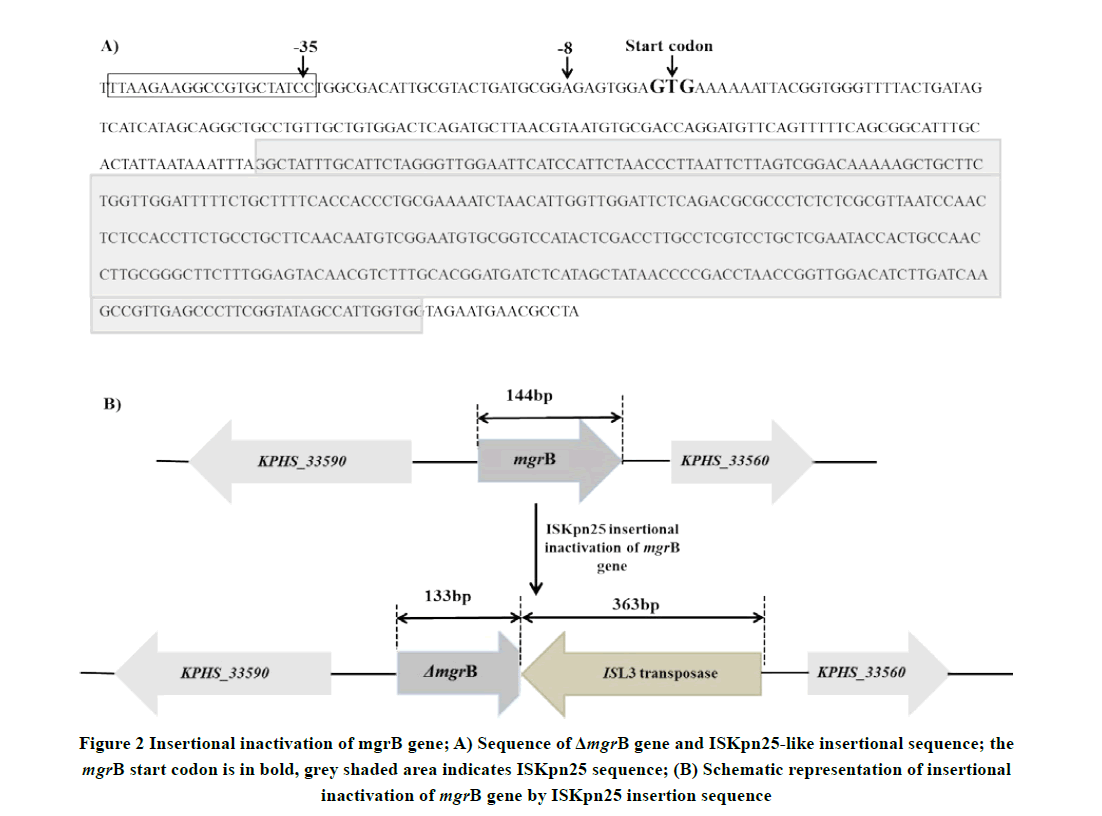
All isolates in the present study contain plasmids. However, PCR amplification of mcr-1 gene was negative for all isolates. PCR amplification targeting mgrB gene was carried out to determine the presence of colistin resistance. Targeted 250 bp PCR amplification gave an amplicon of 510 bp (Figure 1), indicating the presence of an insertion sequence. Sequence analysis of the amplicon revealed that the ISL3 family transposase (ISKpn25 ORF4) of 363 bp targeted in the reverse orientation, downstream of 133 bp of mgrB gene led to insertional inactivation (Figure 2). The NCBI accession number of the seven house-keeping gene sequences used for MLST and the mgrB gene sequence of colistin-resistant carbapenem-resistant K. pneumoniae CKpCR1 were MG367191-MG367198.
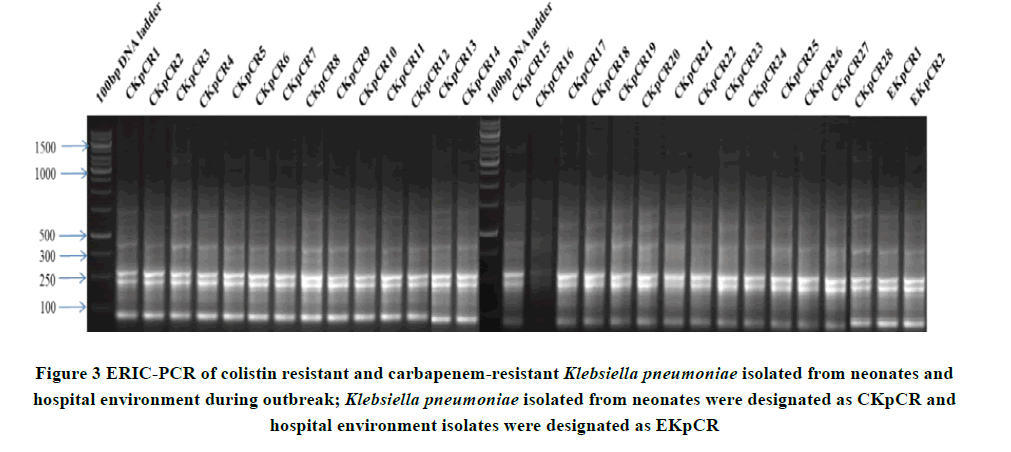
Based on ERIC-PCR band similarity (Figure 3), K. pneumoniae strains were deemed to be representative of a clonal. Multi-locus sequence typing showed that isolates belonged to ST11 with allelic variation 3, 3, 1, 1, 1, 1, 4.
Discussion
In this study, we described and characterized a clonal outbreak of K. pneumoniae bacteremia in the NICU of a tertiary care hospital in the state of Jharkhand, India. All of the neonates in the study exhibited respiratory distress, low platelets associated with diminished spontaneous activity and feeding intolerance, positive blood culture for bacteria with similar cultural characteristics, and antibiotic resistance pattern. The observed mortality was 35.7% over two months. Majhi, et al. have reported detailed clinical parameters during this outbreak, including the severity of illness, risk factors such prematurity, birth asphyxia, babies who had surgery, double volume exchange transfusion, and prior antibiotic usages [18]. This outbreak was caused by a colistin-resistant blaOXA-48 and blaSHV-34 producing K. pneumoniae ST11 that was susceptible only to tigecycline and co-trimoxazole. Proper surveillance and actions like removal of gynecological OT oxygen nozzle and suction, lead to overcome the outbreak.
At present, reserved antibiotic colistin and newly developed glycylcycline antibiotic tigecycline are the few available antibiotics for treating carbapenem-resistant bacteria. Following excessive use, bacteria have progressively developed resistance toward colistin. In 2004, the first clinical outbreak of colistin-resistant K. pneumoniae was reported in Greece [24]. Since then, the emergence of colistin-resistant K. pneumoniae was reported from various parts of the world [25]. Several recent studies highlight the loss-of-function mutations of the mgrB gene responsible for the emergence of colistin resistance in K. pneumoniae. However, other mechanisms, such as the role of efflux pump and plasmid-mediated colistin resistance protein mcr-1 were reported to be associated with colistin resistance in K. pneumoniae [26,27]. In the present study, we found insertional inactivation of the mgrB gene by transposase of ISL3 family that might be responsible for colistin resistance in all the K. pneumoniae isolates. Similar ISL3, a transposable restriction-modification system of 8154 nucleotides inserted in mgrB nucleotide position 133 was reported in colistinresistant K. pneumoniae ST258 isolated from two Italian and eight Greek patients [28]. They reported that ISL3 was located on pKpQIL-like plasmids and may transpose into the chromosome. Single locus variant of ST11, colistin-resistant K. pneumoniae ST258 with the insertional inactivation of mgrB gene by an IS5-like element was reported [15].
In clinical isolates of Klebsiella pneumoniae, colistin resistance has been reported to be mainly associated with the sequence types ST 258, ST 512, and ST147 [29,30]. However, in our study, colistin-resistant K. pneumoniae has been observed in sequence type ST11. In a SMART surveillance program during 2008 and 2009, K. pneumoniae ST11 producing blaOXA-48-like and NDMs were reported from India and many parts of the world [31]. In India, colistin-resistant K. pneumoniae ST11 were reported from a tertiary care hospital carried β-lactam, fluoroquinolones, aminoglycoside, and fosfomycin resistant genes on the genomic DNA [32]. We found that all the isolated K. pneumoniae have ESBL (blaSHV-34) and carbapenemase (blaOXA-48) genes though the fluoroquinolones and aminoglycoside resistance study were not carried out in the present study.
Conclusion
In conclusion, this study showed the emergence of bacterial resistance to the last-resort antibiotic, especially in the intensive care unit of the hospital is of great concern and required judicious use of antibiotics with proper susceptibility analysis as well as continuous surveillance to minimize the emergence of new clones.
Declarations
Conflicts of Interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
References
- World Health Organization. "Antimicrobial resistance global report on surveillance: 2014 summary." No. WHO/HSE/PED/AIP/2014.2. World Health Organization, 2014.
- Podschun, Rainer, and U. Ullmann. "Klebsiella spp. as nosocomial pathogens: Epidemiology, taxonomy, typing methods, and pathogenicity factors." Clinical Microbiology Reviews, Vol. 11, No. 4, 1998, pp. 589-603.
- Victor, L. Yu, et al. "Virulence characteristics of Klebsiella and clinical manifestations of K. pneumoniae bloodstream infections." Emerging Infectious Diseases, Vol. 13, No. 7, 2007, pp. 986-93.
- Boucher, Helen W., et al. "Bad bugs, no drugs: No ESKAPE! An update from the Infectious Diseases Society of America." Clinical Infectious Diseases, Vol. 48, No. 1, 2009, pp. 1-12.
- Sankar, M. Jeeva, et al. "Sepsis in the newborn." The Indian Journal of Pediatrics, Vol. 75, No. 3, 2008, pp. 261-66.
- Borgia, Sergio, et al. "Outbreak of carbapenem-resistant Enterobacteriaceae containing bla NDM-1, Ontario, Canada." Clinical Infectious Diseases, Vol. 55, No. 11, 2012, pp. e109-17.
- Poirel, Laurent, et al. "NDM-1-producing Klebsiella pneumoniae in Mauritius." Antimicrobial Agents and Chemotherapy, Vol. 56, No. 1, 2012, pp. 598-99.
- Teo, J. W. P., et al. "Emergence of clinical Klebsiella pneumoniae producing OXA-232 carbapenemase in Singapore." New Microbes and New Infections, Vol. 1, No. 1, 2013, pp. 13-15.
- Wener, Kenneth M., et al. "Treatment with fluoroquinolones or with β-lactam-β-lactamase inhibitor combinations is a risk factor for isolation of extended-spectrum-β-lactamase-producing Klebsiella species in hospitalized patients." Antimicrobial Agents and Chemotherapy, Vol. 54, No. 5, 2010, pp. 2010-16.
- Li, Jian, et al. "Colistin: The re-emerging antibiotic for multidrug-resistant Gram-negative bacterial infections." The Lancet Infectious Diseases, Vol. 6, No. 9, 2006, pp. 589-601.
- Brink, Adrian J., et al. "Multicomponent antibiotic substances produced by fermentation: Implications for regulatory authorities, critically ill patients and generics." International Journal of Antimicrobial Agents, Vol. 43, No. 1, 2014, pp. 1-6.
- Bergen, Phillip J., et al. "Pharmacokinetics and pharmacodynamics of ‘old’polymyxins: What is new?" Diagnostic Microbiology and Infectious Disease, Vol. 74, No. 3, 2012, pp. 213-23.
- Ah, Young-Mi, Ah-Jung Kim, and Ju-Yeun Lee. "Colistin resistance in Klebsiella pneumoniae." International Journal of Antimicrobial Agents, Vol. 44, No. 1, 2014, pp. 8-15.
- Singh, Santosh Kumar, et al. "Efflux mediated colistin resistance in diverse clones of Klebsiella pneumoniae from aquatic environment." Microbial Pathogenesis, Vol. 102, 2017, pp. 109-12.
- Cannatelli, Antonio, et al. "In vivo emergence of colistin resistance in Klebsiella pneumoniae producing KPC-type carbapenemases mediated by insertional inactivation of the PhoQ/PhoP mgrB regulator." Antimicrobial Agents and Chemotherapy, Vol. 57, No. 11, 2013, pp. 5521-26.
- Cheng, Yi-Hsiang, et al. "Colistin resistance mechanisms in Klebsiella pneumoniae strains from Taiwan." Antimicrobial Agents and Chemotherapy, Vol. 59, No. 5, 2015, pp. 2909-13.
- Liu, Yi-Yun, et al. "Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study." The Lancet Infectious Diseases, Vol. 16 No. 2, 2016, pp. 161-68.
- Majhi, Jatin Kumar, Asit Kumar Mishra, and Arati Behera. "A study on outbreak of carbapenem plus colistin-resistant Klebsiella pneumoniae sepsis in newborns in a nursery of a tertiary care hospital." Journal of Evidence Based Medicine and Healthcare, Vol. 4, 2017, pp. 119-22.
- Jorgensen, James H., and John D. Turnidge. "Susceptibility test methods: Dilution and disk diffusion methods." Manual of Clinical Microbiology, 2015, pp. 1253-73.
- Patel, Jean B., F. R. Cockerill, and Patricia A. Bradford. "Performance standards for antimicrobial susceptibility testing: Twenty-fifth informational supplement." Clinical and Laboratory Standards Institute, Vol. 35, No. 3, 2015, pp. 29-50.
- Yilmaz, N. O., et al. "Detection of plasmid-mediated AmpC β-lactamase in Escherichia coli and Klebsiella pneumoniae." Indian Journal of Medical Microbiology, Vol. 31, No. 1, 2013, pp. 53-59.
- Munoz, Marcos A., et al. "Molecular epidemiology of two Klebsiella pneumoniae mastitis outbreaks on a dairy farm in New York State." Journal of Clinical Microbiology, Vol. 45, No. 12, 2007, pp. 3964-71.
- Diancourt, Laure, et al. "Multilocus sequence typing of Klebsiella pneumoniae nosocomial isolates." Journal of Clinical Microbiology, Vol. 43, No. 8, 2005, pp. 4178-82.
- Antoniadou, Anastasia, et al. "Colistin-resistant isolates of Klebsiella pneumoniae emerging in intensive care unit patients: First report of a multiclonal cluster." Journal of Antimicrobial Chemotherapy, Vol. 59, No. 4, 2007, pp. 786-90.
- Olaitan, Abiola Olumuyiwa, et al. "Worldwide emergence of colistin resistance in Klebsiella pneumoniae from healthy humans and patients in Lao PDR, Thailand, Israel, Nigeria and France owing to inactivation of the PhoP/PhoQ regulator mgrB: An epidemiological and molecular study." International Journal of Antimicrobial Agents, Vol. 44, No. 6, 2014, pp. 500-07.
- Singh, Santosh Kumar, et al. "Efflux mediated colistin resistance in diverse clones of Klebsiella pneumoniae from aquatic environment." Microbial Pathogenesis, Vol. 102, 2017, pp. 109-12.
- Gu, Dan-xia, et al. "Detection of colistin resistance gene mcr-1 in hypervirulent Klebsiella pneumoniae and Escherichia coli isolates from an infant with diarrhea in China." Antimicrobial Agents and Chemotherapy, Vol. 60, No. 8, 2016, pp. 5099-100.
- Giordano, Cesira, et al. "Expansion of KPC-producing Klebsiella pneumoniae with various mgrB mutations giving rise to colistin resistance: The role of ISL3 on plasmids." International Journal of Antimicrobial Agents, Vol. 51, No. 2, 2018, pp. 260-65.
- Cannatelli, Antonio, et al. "In vivo evolution to colistin resistance by PmrB sensor kinase mutation in KPC-producing Klebsiella pneumoniae is associated with low-dosage colistin treatment." Antimicrobial Agents and Chemotherapy, Vol. 58, No. 8, 2014, pp. 4399-403.
- Mammina, C., et al. "Ongoing spread of colistin-resistant Klebsiella pneumoniae in different wards of an acute general hospital, Italy, June to December 2011." Eurosurveillance, Vol. 17, No. 33, 2012, p. 20248.
- Lascols, Christine, et al. "Surveillance and molecular epidemiology of Klebsiella pneumoniae isolates that produce carbapenemases: First report of OXA-48-like enzymes in North America." Antimicrobial Agents and Chemotherapy, Vol. 57, No. 1, 2013, pp. 130-36.
- Mathur, Purva, et al. "First report on a cluster of colistin-resistant Klebsiella pneumoniae strains isolated from a tertiary care center in India: Whole-genome shotgun sequencing." Genome Announcements, Vol. 5, No. 5, 2017, pp. e01466-16.